

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
null
SMILES: CCCC(NC(=O)[C@@H]1[C@@H]2[C@H](CN1C(=O)[C@@H](NC(=O)OC(C)(C)C)C1CCCCC1)C2(C)C)C(=O)C(=O)NCC(=O)NCc1ccccc1
InChI Key: InChIKey=FQWQFMJDELOUET-JFLFQVESSA-N
PDB links: 1 PDB ID contains inhibitors having a similarity of 90% to this monomer.
| Target/Host (Institution) | Ligand | Target/Host Links | Ligand Links | Trg + Lig Links | Ki nM | ΔG° kcal/mole | IC50 nM | Kd nM | EC50/IC50 nM | koff s-1 | kon M-1s-1 | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Genome polyprotein (Hepatitis C virus (HCV genotype 1a, isolate H)) | BDBM12267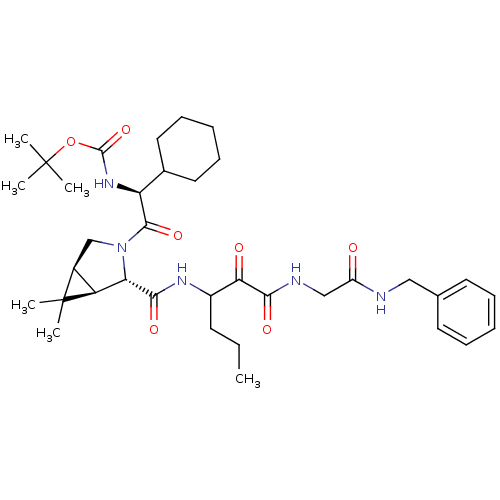 (1,1-Dimethylethyl-[1(S)-cyclohexyl-2-[(1R,5S)-6,6-...) | PDB MMDB UniProtKB/SwissProt B.MOAD DrugBank GoogleScholar AffyNet | PC cid PC sid UniChem Patents Similars | Article PubMed | 56 | -10.1 | n/a | n/a | n/a | n/a | n/a | 6.5 | 30 |
Schering-Plough Research Institute | Assay Description Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I... | J Med Chem 49: 6074-86 (2006) Article DOI: 10.1021/jm060325b BindingDB Entry DOI: 10.7270/Q2DF6PF6 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||