

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
BDBM190659 PT156
SMILES: CCCCCCc1cc(=O)c(Oc2ccc(cc2)[N+]([O-])=O)cn1C
InChI Key: InChIKey=SRHFLDROUHFSKC-UHFFFAOYSA-N
| Target/Host (Institution) | Ligand | Target/Host Links | Ligand Links | Trg + Lig Links | Ki nM | ΔG° kcal/mole | IC50 nM | Kd nM | EC50/IC50 nM | koff s-1 | kon M-1s-1 | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Enoyl-[acyl-carrier-protein] reductase [NADH] (Yersinia pestis (Enterobacteria)) | BDBM190659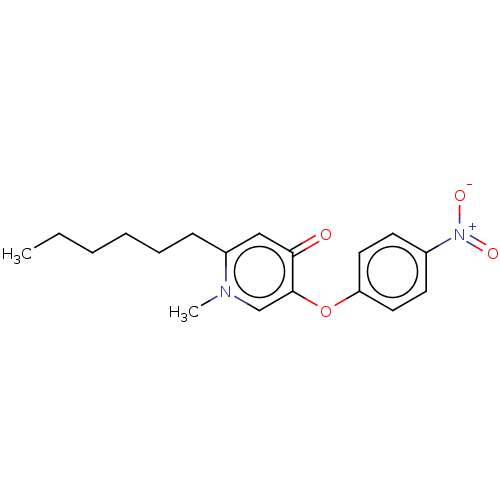 (PT156) | PDB UniProtKB/SwissProt UniProtKB/TrEMBL GoogleScholar AffyNet | PC cid PC sid UniChem Similars | Article PubMed | 200 | -9.13 | n/a | n/a | n/a | n/a | n/a | 8.0 | 25 |
University of Würzburg | Assay Description Steady-state kinetics were performed on a Cary 100 Bio (Varian) spectrometer at 25 °C using 30 mM PIPES, 150 mM NaCl, and 1.0 mM EDTA (pH 8.0) as... | Biochemistry 55: 2992-3006 (2016) Article DOI: 10.1021/acs.biochem.5b01301 BindingDB Entry DOI: 10.7270/Q25D8QM3 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Enoyl-[acyl-carrier-protein] reductase [NADH] [T276S] (Yersinia pestis (Enterobacteria)) | BDBM190659 (PT156) | PDB GoogleScholar AffyNet | PC cid PC sid UniChem Similars | Article PubMed | 500 | -8.59 | n/a | n/a | n/a | n/a | n/a | 8.0 | 25 |
University of Würzburg | Assay Description Steady-state kinetics were performed on a Cary 100 Bio (Varian) spectrometer at 25 °C using 30 mM PIPES, 150 mM NaCl, and 1.0 mM EDTA (pH 8.0) as... | Biochemistry 55: 2992-3006 (2016) Article DOI: 10.1021/acs.biochem.5b01301 BindingDB Entry DOI: 10.7270/Q25D8QM3 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Enoyl-[acyl-carrier-protein] reductase [NADH] (Yersinia pestis (Enterobacteria)) | BDBM190659 (PT156) | PDB UniProtKB/SwissProt UniProtKB/TrEMBL GoogleScholar AffyNet | PC cid PC sid UniChem Similars | Article PubMed | n/a | n/a | 200 | n/a | n/a | n/a | n/a | n/a | n/a |
University of Würzburg | Assay Description IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet... | Biochemistry 55: 2992-3006 (2016) Article DOI: 10.1021/acs.biochem.5b01301 BindingDB Entry DOI: 10.7270/Q25D8QM3 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Enoyl-[acyl-carrier-protein] reductase [NADH] [T276S] (Yersinia pestis (Enterobacteria)) | BDBM190659 (PT156) | PDB GoogleScholar AffyNet | PC cid PC sid UniChem Similars | Article PubMed | n/a | n/a | 1.00E+3 | n/a | n/a | n/a | n/a | n/a | n/a |
University of Würzburg | Assay Description IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet... | Biochemistry 55: 2992-3006 (2016) Article DOI: 10.1021/acs.biochem.5b01301 BindingDB Entry DOI: 10.7270/Q25D8QM3 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||