

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
null
SMILES: [#6]-[#8]-[#6@@H]1-[#6@@H](-[#8]-[#6](-[#7])=O)-[#6@@H](-[#8])-[#6@H](-[#8]-c2ccc3c(-[#8])c(-[#7]-[#6](=O)-c4ccc(-[#8])c(-[#6]\[#6]=[#6](\[#6])-[#6])c4)c(=O)oc3c2-[#6])-[#8]C1([#6])[#6]
InChI Key: InChIKey=YJQPYGGHQPGBLI-KGSXXDOSSA-N
PDB links: 11 PDB IDs match this monomer.
| Target/Host (Institution) | Ligand | Target/Host Links | Ligand Links | Trg + Lig Links | Ki nM | ΔG° kcal/mole | IC50 nM | Kd nM | EC50/IC50 nM | koff s-1 | kon M-1s-1 | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| DNA gyrase subunit A/B (Staphylococcus aureus) | BDBM282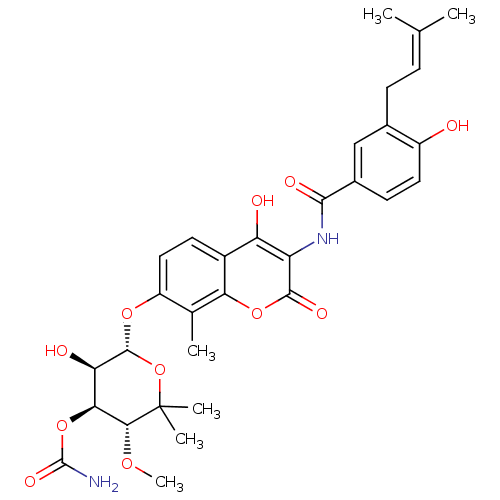 ((3R,4S,5R,6R)-5-hydroxy-6-[(2-hydroxy-3-{[4-hydrox...) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD DrugBank GoogleScholar AffyNet | Purchase DrugBank KEGG MMDB PC cid PC sid PDB UniChem Similars | DrugBank PDB Article PubMed | 10 | -11.1 | n/a | n/a | n/a | n/a | n/a | 7.5 | 30 |
Vertex | Assay Description Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ... | J Med Chem 51: 5243-63 (2008) Article DOI: 10.1021/jm800318d BindingDB Entry DOI: 10.7270/Q2J67F7T | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| DNA gyrase subunit A/B (Escherichia coli (strain K12)) | BDBM282 ((3R,4S,5R,6R)-5-hydroxy-6-[(2-hydroxy-3-{[4-hydrox...) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD DrugBank GoogleScholar AffyNet | Purchase DrugBank KEGG MMDB PC cid PC sid PDB UniChem Similars | Article PubMed | 14 | n/a | n/a | n/a | n/a | n/a | n/a | n/a | n/a |
Vertex | Assay Description Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ... | J Med Chem 51: 5243-63 (2008) Article DOI: 10.1021/jm800318d BindingDB Entry DOI: 10.7270/Q2J67F7T | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| DNA topoisomerase 4 subunit A/B (Escherichia coli (strain K12)) | BDBM282 ((3R,4S,5R,6R)-5-hydroxy-6-[(2-hydroxy-3-{[4-hydrox...) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD DrugBank GoogleScholar AffyNet | Purchase DrugBank KEGG MMDB PC cid PC sid PDB UniChem Similars | PDB Article PubMed | 110 | n/a | n/a | n/a | n/a | n/a | n/a | n/a | n/a |
Vertex | Assay Description Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ... | J Med Chem 51: 5243-63 (2008) Article DOI: 10.1021/jm800318d BindingDB Entry DOI: 10.7270/Q2J67F7T | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| DNA gyrase subunit B (Staphylococcus aureus) | BDBM282 ((3R,4S,5R,6R)-5-hydroxy-6-[(2-hydroxy-3-{[4-hydrox...) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD DrugBank GoogleScholar AffyNet | Purchase DrugBank KEGG MMDB PC cid PC sid PDB UniChem Similars | DrugBank PDB Article PubMed | n/a | n/a | 30 | n/a | n/a | n/a | n/a | 7.6 | 25 |
Birla Institute of Technology & Science-Pilani, Hyderabad Campus | Assay Description Supercoiling assay was performed using the commercially available kit (DNA gyrase supercoiling assay kit: SAS4001) from Inspiralis Pvt.limited, Norwi... | Chem Biol Drug Des 86: 918-25 (2015) Article DOI: 10.1111/cbdd.12529 BindingDB Entry DOI: 10.7270/Q2610Z29 | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| DNA gyrase subunit B (Staphylococcus aureus) | BDBM282 ((3R,4S,5R,6R)-5-hydroxy-6-[(2-hydroxy-3-{[4-hydrox...) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD DrugBank GoogleScholar AffyNet | Purchase DrugBank KEGG MMDB PC cid PC sid PDB UniChem Similars | DrugBank PDB Article PubMed | n/a | n/a | 125 | n/a | n/a | n/a | n/a | 7.5 | n/a |
Birla Institute of Technology & Science-Pilani, Hyderabad Campus | Assay Description Initially, equimolar quantities of about 0.5�M each of E. coli gyraseA and S. aureus gyraseB were incubated for a period of 45 min with salmon sperm ... | Chem Biol Drug Des 86: 918-25 (2015) Article DOI: 10.1111/cbdd.12529 BindingDB Entry DOI: 10.7270/Q2610Z29 | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| DNA gyrase subunit B (Mycobacterium smegmatis) | BDBM282 ((3R,4S,5R,6R)-5-hydroxy-6-[(2-hydroxy-3-{[4-hydrox...) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD GoogleScholar AffyNet | Purchase DrugBank KEGG MMDB PC cid PC sid PDB UniChem Similars | PDB Article PubMed | n/a | n/a | 180 | n/a | n/a | n/a | n/a | n/a | n/a |
Birla Institute of Technology& Science-Pilani Curated by ChEMBL | Assay Description Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate... | Bioorg Med Chem 23: 2062-78 (2015) Article DOI: 10.1016/j.bmc.2015.03.004 BindingDB Entry DOI: 10.7270/Q2HT2R1V | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| Efflux pump membrane transporter (Escherichia coli) | BDBM282 ((3R,4S,5R,6R)-5-hydroxy-6-[(2-hydroxy-3-{[4-hydrox...) | PDB MMDB KEGG UniProtKB/TrEMBL B.MOAD DrugBank GoogleScholar AffyNet | Purchase DrugBank KEGG MMDB PC cid PC sid PDB UniChem Similars | Article PubMed | n/a | n/a | n/a | 5.30E+5 | n/a | n/a | n/a | 7.5 | n/a |
University of Oklahoma | Assay Description Binding of inhibitors and substrates to immobilized AcrB. | Chem Biol 18: 454-63 (2011) Article DOI: 10.1016/j.chembiol.2011.02.011 BindingDB Entry DOI: 10.7270/Q2DN43JT | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Cell (A) | Syringe (B) | Cell Links | Syringe Links | Cell + Syr Links | ΔG° kcal/mole | -TΔS° kcal/mole | ΔH° kcal/mole | log K | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|
| DNA Gyrase (Escherichia coli) | BDBM282 ((3R,4S,5R,6R)-5-hydroxy-6-[(2-hydroxy-3-{[4-hydrox...) | GoogleScholar PDB | CHEBI DrugBank KEGG MMDB PC cid PC sid PDB | -10.2 | 2.17 | -12.4 | 7.45 | 7.5 | 25 | |
UMR CNRS 6032 | Biochemistry 41: 7217-23 (2002) | |||||||||