

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
BDBM36343 CID24762173::N2-trans-(4-Amino-cyclohexyl)-N6-(4-morpholin-4-ylmethyl-phenyl)-9-phenyl-9H-purine-2,6-diamine, 11
SMILES: N[C@H]1CC[C@@H](CC1)Nc1nc(Nc2ccc(CN3CCOCC3)cc2)c2ncn(-c3ccccc3)c2n1
InChI Key: InChIKey=WXTHNIOSZSQDJR-AFARHQOCSA-N
| Target/Host (Institution) | Ligand | Target/Host Links | Ligand Links | Trg + Lig Links | Ki nM | ΔG° kcal/mole | IC50 nM | Kd nM | EC50/IC50 nM | koff s-1 | kon M-1s-1 | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Calcium-dependent protein kinase 1 (Plasmodium Falciparum) | BDBM36343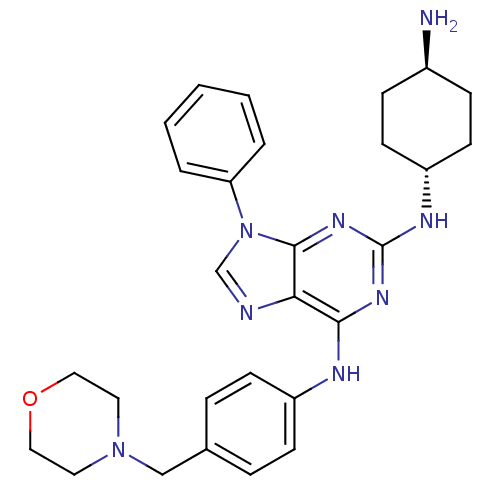 (CID24762173 | N2-trans-(4-Amino-cyclohexyl)-N6-(4-...) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD GoogleScholar AffyNet | PC cid PC sid UniChem Similars | Article PubMed | n/a | n/a | n/a | n/a | 174 | n/a | n/a | 7.5 | 24 |
The Scripps Research Institute | Nat Chem Biol 4: 347-56 (2008) Article DOI: 10.1038/nchembio.87 BindingDB Entry DOI: 10.7270/Q2TB1573 | ||||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Calcium-dependent protein kinase 1 (Plasmodium Falciparum) | BDBM36343 (CID24762173 | N2-trans-(4-Amino-cyclohexyl)-N6-(4-...) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD GoogleScholar AffyNet | PC cid PC sid UniChem Similars | Article PubMed | n/a | n/a | 703 | n/a | n/a | n/a | n/a | 7.5 | 24 |
The Scripps Research Institute | Assay Description The test compounds and enzyme were mixed in buffer, and the substrate and [gamma-33P] ATP/ATP were added to the mixture to initiate the reaction. Aft... | Nat Chem Biol 4: 347-56 (2008) Article DOI: 10.1038/nchembio.87 BindingDB Entry DOI: 10.7270/Q2TB1573 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||