

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
BDBM50049379 (S)-3-[(S)-2-({1-[(S)-2-Amino-3-(4-hydroxy-phenyl)-propionylamino]-cyclohexanecarbonyl}-amino)-3-phenyl-propionylamino]-N-{(S)-1-[(S)-1-(carbamoylmethyl-carbamoyl)-2-methyl-propylcarbamoyl]-2-methyl-propyl}-succinamic acid::CHEMBL165263
SMILES: CC(C)[C@H](NC(=O)[C@@H](NC(=O)[C@H](CC(O)=O)NC(=O)[C@H](Cc1ccccc1)NC(=O)C1(CCCCC1)NC(=O)[C@@H](N)Cc1ccc(O)cc1)C(C)C)C(=O)NCC(N)=O
InChI Key: InChIKey=UEVFYKFKEUHPLB-WDEDYMKJSA-N
Data: 2 KI
| Target/Host (Institution) | Ligand | Target/Host Links | Ligand Links | Trg + Lig Links | Ki nM | ΔG° kcal/mole | IC50 nM | Kd nM | EC50/IC50 nM | koff s-1 | kon M-1s-1 | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Opioid receptors; mu & delta (Rattus norvegicus (rat)) | BDBM50049379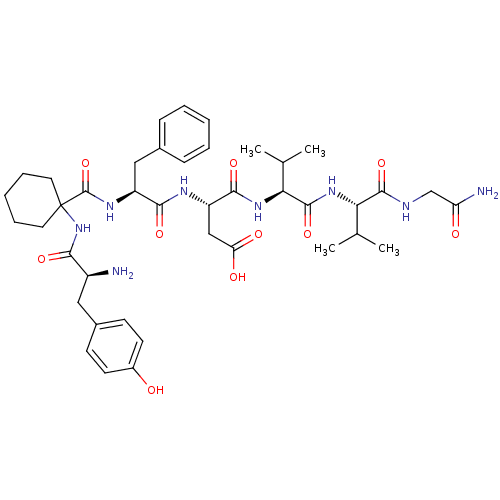 ((S)-3-[(S)-2-({1-[(S)-2-Amino-3-(4-hydroxy-phenyl)...) | PDB Reactome pathway KEGG UniProtKB/SwissProt B.MOAD GoogleScholar AffyNet | CHEMBL PC cid PC sid UniChem Similars | Article PubMed | 2.48 | n/a | n/a | n/a | n/a | n/a | n/a | n/a | n/a |
University of Ferrara Curated by ChEMBL | Assay Description Displacement of [3H]-DPDPE (0.63 nM) from Opioid receptor delta 1 | J Med Chem 39: 773-80 (1996) Article DOI: 10.1021/jm950490j BindingDB Entry DOI: 10.7270/Q2QV3KKJ | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Mu-type opioid receptor (Rattus norvegicus (rat)) | BDBM50049379 ((S)-3-[(S)-2-({1-[(S)-2-Amino-3-(4-hydroxy-phenyl)...) | PDB Reactome pathway KEGG UniProtKB/SwissProt B.MOAD GoogleScholar AffyNet | CHEMBL PC cid PC sid UniChem Similars | Article PubMed | 3.11E+3 | n/a | n/a | n/a | n/a | n/a | n/a | n/a | n/a |
University of Ferrara Curated by ChEMBL | Assay Description Displacement of [3H]-DAGO (1.28 nM) from Opioid receptor mu 1 | J Med Chem 39: 773-80 (1996) Article DOI: 10.1021/jm950490j BindingDB Entry DOI: 10.7270/Q2QV3KKJ | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||