

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Target/Host (Institution) | Ligand | Target/Host Links | Ligand Links | Trg + Lig Links | Ki nM | ΔG° kJ/mole | IC50 nM | Kd nM | EC50/IC50 nM | koff s-1 | kon M-1s-1 | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| DNA gyrase subunit B (205/205 = 100%)† (Escherichia coli (strain K12)) | BDBM50330317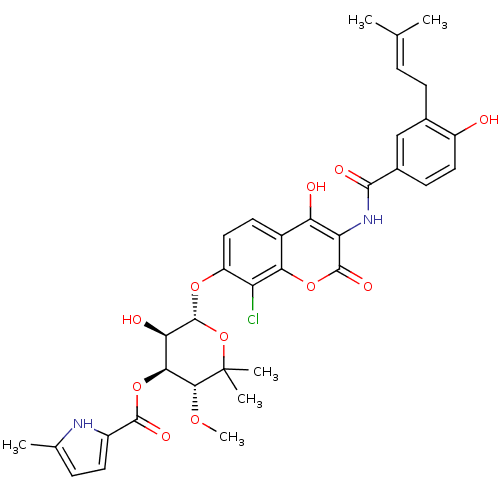 (5-Methyl-1H-pyrrole-2-carboxylic acid (3R,4S,5R,6S...) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD DrugBank GoogleScholar AffyNet | Purchase CHEMBL MCE KEGG MMDB PC cid PC sid PDB UniChem Similars | MMDB PDB Article PubMed | n/a | n/a | 73 | n/a | n/a | n/a | n/a | n/a | n/a |
Universität Tübingen Curated by ChEMBL | Assay Description Inhibition of Escherichia coli JM109 DNA gyrase subunit B assessed as ATP hydrolysis by ATPase assay | Antimicrob Agents Chemother 52: 1982-90 (2008) Article DOI: 10.1128/AAC.01235-07 BindingDB Entry DOI: 10.7270/Q2PG1S0J | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| DNA gyrase subunit B (205/205 = 100%)† (Escherichia coli) | BDBM50330317 (5-Methyl-1H-pyrrole-2-carboxylic acid (3R,4S,5R,6S...) | PDB MMDB UniProtKB/TrEMBL B.MOAD GoogleScholar AffyNet | Purchase CHEMBL MCE KEGG MMDB PC cid PC sid PDB UniChem Similars | MMDB PDB Article PubMed | n/a | n/a | n/a | 1.20 | n/a | n/a | n/a | n/a | n/a |
University of Ljubljana Curated by ChEMBL | Assay Description Binding affinity to Escherichia coli GyrB | J Med Chem 54: 915-29 (2011) Article DOI: 10.1021/jm101121s BindingDB Entry DOI: 10.7270/Q2NS0W23 | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| Cell (A) | Syringe (B) | Cell Links | Syringe Links | Cell + Syr Links | ΔG° kJ/mole | -TΔS° kJ/mole | ΔH° kJ/mole | log K | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|
| DNA Gyrase (205/205 = 100%)† (Escherichia coli) | BDBM283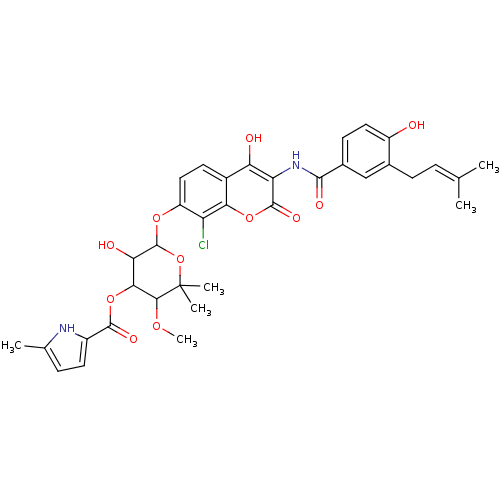 (6-[(8-chloro-4-hydroxy-3-{[4-hydroxy-3-(3-methylbu...) | GoogleScholar PDB | PC cid PC sid PDB | -51 | -11.4 | -39.6 | 8.93 | 7.5 | 25 | |
UMR CNRS 6032 | Biochemistry 41: 7217-23 (2002) | |||||||||
| Cell (A) | Syringe (B) | Cell Links | Syringe Links | Cell + Syr Links | ΔG° kJ/mole | -TΔS° kJ/mole | ΔH° kJ/mole | log K | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|
| DNA Gyrase (205/205 = 100%)† (Escherichia coli) | BDBM283 (6-[(8-chloro-4-hydroxy-3-{[4-hydroxy-3-(3-methylbu...) | GoogleScholar PDB | PC cid PC sid PDB | -51 | -11.4 | -39.6 | 8.93 | 7.5 | 25 | |
UMR CNRS 6032 | Biochemistry 41: 7217-23 (2002) | |||||||||