 Found 3 hits Enzyme Inhibition Constant Data
Found 3 hits Enzyme Inhibition Constant Data Target/Host
(Institution) | Ligand | Target/Host
Links | Ligand
Links | Trg + Lig
Links | Ki
nM | ΔG°
kJ/mole | IC50
nM | Kd
nM | EC50/IC50
nM | koff
s-1 | kon
M-1s-1 | pH | Temp
°C |
|---|
ATP-dependent molecular chaperone HSP82
(214/214 = 100%)†
(Saccharomyces cerevisiae) | BDBM68260
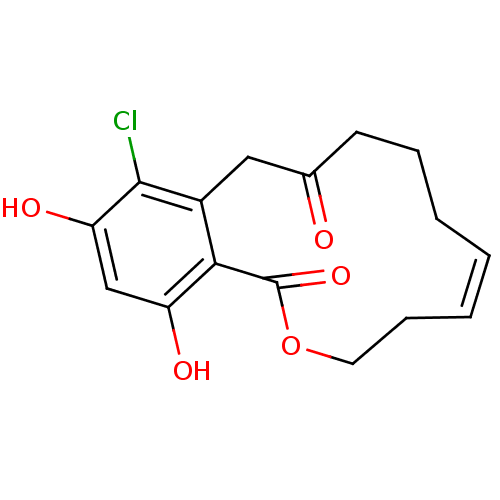
(Microlactone, 15f cis)Show SMILES Oc1cc(O)c2c(CC(=O)CCC\C=C/CCOC2=O)c1Cl |c:13| Show InChI InChI=1S/C16H17ClO5/c17-15-11-8-10(18)6-4-2-1-3-5-7-22-16(21)14(11)12(19)9-13(15)20/h1,3,9,19-20H,2,4-8H2/b3-1- | PDB
MMDB
Reactome pathway
KEGG
UniProtKB/SwissProt
B.MOAD
DrugBank
GoogleScholar
AffyNet 
| DrugBank
MMDB
PC cid
PC sid
PDB
UniChem
Similars
| MMDB
PDB
Article
PubMed
| n/a | n/a | 500 | n/a | n/a | n/a | n/a | n/a | n/a |
University of Nottingham
| Assay Description
Binding competition assay with a fluorescent probe. |
Chem Biol 13: 1203-15 (2006)
Article DOI: 10.1016/j.chembiol.2006.09.015
BindingDB Entry DOI: 10.7270/Q2930RMM |
More data for this
Ligand-Target Pair | 
3D Structure (crystal) |
ATP-dependent molecular chaperone HSP82
(214/214 = 100%)†
(Saccharomyces cerevisiae) | BDBM68260

(Microlactone, 15f cis)Show SMILES Oc1cc(O)c2c(CC(=O)CCC\C=C/CCOC2=O)c1Cl |c:13| Show InChI InChI=1S/C16H17ClO5/c17-15-11-8-10(18)6-4-2-1-3-5-7-22-16(21)14(11)12(19)9-13(15)20/h1,3,9,19-20H,2,4-8H2/b3-1- | PDB
MMDB
Reactome pathway
KEGG
UniProtKB/SwissProt
B.MOAD
DrugBank
GoogleScholar
AffyNet 
| DrugBank
MMDB
PC cid
PC sid
PDB
UniChem
Similars
| MMDB
PDB
Article
PubMed
| n/a | n/a | 850 | n/a | n/a | n/a | n/a | n/a | n/a |
University of Nottingham
| Assay Description
Binding competition assay with a fluorescent probe. |
Chem Biol 13: 1203-15 (2006)
Article DOI: 10.1016/j.chembiol.2006.09.015
BindingDB Entry DOI: 10.7270/Q2930RMM |
More data for this
Ligand-Target Pair | 
3D Structure (crystal) |
ATP-dependent molecular chaperone HSP82
(214/214 = 100%)†
(Saccharomyces cerevisiae) | BDBM68260

(Microlactone, 15f cis)Show SMILES Oc1cc(O)c2c(CC(=O)CCC\C=C/CCOC2=O)c1Cl |c:13| Show InChI InChI=1S/C16H17ClO5/c17-15-11-8-10(18)6-4-2-1-3-5-7-22-16(21)14(11)12(19)9-13(15)20/h1,3,9,19-20H,2,4-8H2/b3-1- | PDB
MMDB
Reactome pathway
KEGG
UniProtKB/SwissProt
B.MOAD
DrugBank
GoogleScholar
AffyNet 
| DrugBank
MMDB
PC cid
PC sid
PDB
UniChem
Similars
| MMDB
PDB
Article
PubMed
| n/a | n/a | 2.40E+3 | n/a | n/a | n/a | n/a | n/a | n/a |
University of Nottingham
| Assay Description
A colorimetric assay for the release of inorganic phosphate upon hydrolysis of ATP was used to determine the potency of Hsp 90 inhibitor against enzy... |
Chem Biol 13: 1203-15 (2006)
Article DOI: 10.1016/j.chembiol.2006.09.015
BindingDB Entry DOI: 10.7270/Q2930RMM |
More data for this
Ligand-Target Pair | 
3D Structure (crystal) |


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data