

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Target/Host (Institution) | Ligand | Target/Host Links | Ligand Links | Trg + Lig Links | Ki nM | ΔG° kJ/mole | IC50 nM | Kd nM | EC50/IC50 nM | koff s-1 | kon M-1s-1 | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Peptide deformylase (203/203 = 100%)† (Streptococcus pneumoniae (strain ATCC BAA-255 / R6...) | BDBM92760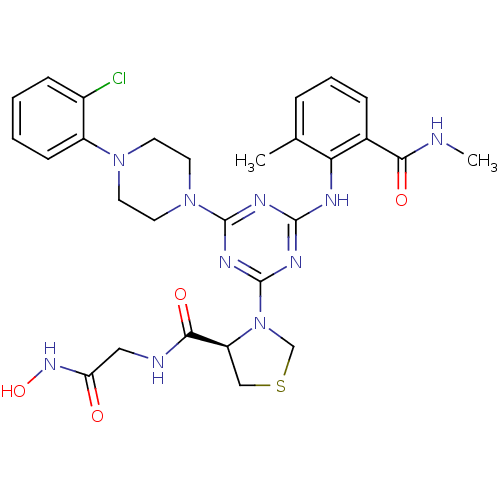 (PDF inhibitor, compound 2) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD DrugBank GoogleScholar AffyNet | MMDB PC cid PC sid PDB UniChem Similars | PDB Article PubMed | 334 | n/a | n/a | n/a | n/a | n/a | n/a | n/a | n/a |
GlaxoSmithKline | Assay Description Binding assay using PDF inhibitors. | Biochemistry 50: 6642-54 (2011) Article DOI: 10.1021/bi200655g BindingDB Entry DOI: 10.7270/Q2MS3RBV | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||