PDB code 3UUO
Identical Ligands in BindingDB
 Found 2 hits Enzyme Inhibition Constant Data
Found 2 hits Enzyme Inhibition Constant Data Target/Host
(Institution) | Ligand | Target/Host
Links | Ligand
Links | Trg + Lig
Links | Ki
nM | ΔG°
kJ/mole | IC50
nM | Kd
nM | EC50/IC50
nM | koff
s-1 | kon
M-1s-1 | pH | Temp
°C |
|---|
cAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A
(329/332 > 99%)†
(Homo sapiens (Human)) | BDBM50362709
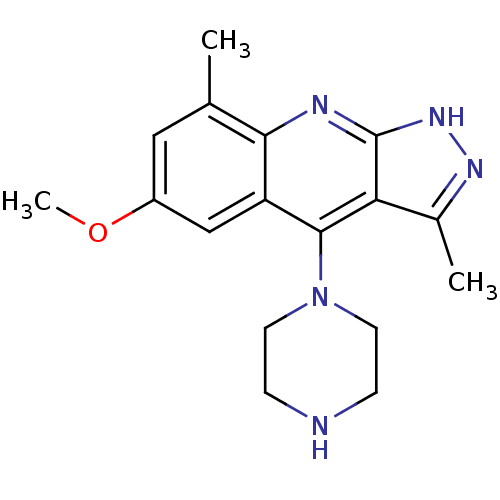
(CHEMBL1939782)Show InChI InChI=1S/C17H21N5O/c1-10-8-12(23-3)9-13-15(10)19-17-14(11(2)20-21-17)16(13)22-6-4-18-5-7-22/h8-9,18H,4-7H2,1-3H3,(H,19,20,21) | PDB
MMDB
Reactome pathway
KEGG
UniProtKB/SwissProt
B.MOAD
DrugBank
antibodypedia
GoogleScholar
AffyNet 
| CHEMBL
MMDB
PC cid
PC sid
PDB
UniChem
Similars
| PDB
Article
PubMed
| 11 | n/a | n/a | n/a | n/a | n/a | n/a | n/a | n/a |
Merck Research Laboratories
Curated by ChEMBL
| Assay Description
Inhibition of recombinant human PDE10A using [3H]cAMP as substrate by scintillation proximity assay |
Bioorg Med Chem Lett 22: 1019-22 (2012)
Article DOI: 10.1016/j.bmcl.2011.11.127
BindingDB Entry DOI: 10.7270/Q2BV7H3J |
More data for this
Ligand-Target Pair | 
3D Structure (crystal) |
cAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A
(329/332 > 99%)†
(Homo sapiens (Human)) | BDBM50362709

(CHEMBL1939782)Show InChI InChI=1S/C17H21N5O/c1-10-8-12(23-3)9-13-15(10)19-17-14(11(2)20-21-17)16(13)22-6-4-18-5-7-22/h8-9,18H,4-7H2,1-3H3,(H,19,20,21) | PDB
MMDB
Reactome pathway
KEGG
UniProtKB/SwissProt
B.MOAD
DrugBank
antibodypedia
GoogleScholar
AffyNet 
| CHEMBL
MMDB
PC cid
PC sid
PDB
UniChem
Similars
| PDB
Article
PubMed
| n/a | n/a | 858 | n/a | n/a | n/a | n/a | n/a | n/a |
Merck Research Laboratories
Curated by ChEMBL
| Assay Description
Inhibition of recombinant human PDE10A using [3H]cAMP as substrate by scintillation proximity assay |
Bioorg Med Chem Lett 22: 1019-22 (2012)
Article DOI: 10.1016/j.bmcl.2011.11.127
BindingDB Entry DOI: 10.7270/Q2BV7H3J |
More data for this
Ligand-Target Pair | 
3D Structure (crystal) |
- Reorder this table by ec50
- Reorder this table by ic50
- Reorder this table by Kd
- Reorder this table by pH
- Reorder this table by T
- Reorder this table by ΔH°
- Reorder this table by -TΔS°
- Change energy units to: kcal/mole
Search BindingMOAD for More Affinity Data:
* indicates data uncertainty>20%
* 0.9 Tanimoto similarity
† Identities from BLAST output



 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data