

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Cell Reactant: | DNA Gyrase | ||
| Syringe Reactant: | BDBM283 | ||
| Meas. Tech.: | Isothermal Titration Calorimetry | ||
| Entry Date: | 03/31/03 | ||
| ΔG°: | -51±0.600 (kJ/mole) | ||
| pH: | 7.5±n/a | ||
| Log10Kb: | 8.93± 8.34 | ||
| Temperature: | 298.15±n/a (K) | ||
| ΔH° : | -39.6±2.40 (kJ/mole) | ||
| ΔHobs : | -39.6±2.40 (kJ/mole) | ||
| Ionic Strength: | n/a | ||
| not known | |||
| Protons Released: | 0 | ||
| ΔCp : | -1.46±0.14 (kJ/mole) | ||
| Stoich. Param.: | 1 | ||
| ΔS° : | 0.0383±0.00340 (kJ/mole-K) | ||
| Comments: | Displacement titration measurements were used because the constant of association was too high to measure directly. | ||
| Citation |  Lafitte, D; Lamour, V; Tsvetkov, PO; Makarov, AA; Klich, M; Deprez, P; Moras, D; Briand, C; Gilli, R DNA gyrase interaction with coumarin-based inhibitors: the role of the hydroxybenzoate isopentenyl moiety and the 5'-methyl group of the noviose. Biochemistry41:7217-23 (2002) [PubMed] Article Lafitte, D; Lamour, V; Tsvetkov, PO; Makarov, AA; Klich, M; Deprez, P; Moras, D; Briand, C; Gilli, R DNA gyrase interaction with coumarin-based inhibitors: the role of the hydroxybenzoate isopentenyl moiety and the 5'-methyl group of the noviose. Biochemistry41:7217-23 (2002) [PubMed] Article | ||
| More Info.: | Get all data from this article , ITC RUN data , Solution Info , Data Fit Method , Instrument Info | ||
| DNA Gyrase | |||
| Source: | Bacterial clone | ||
| Purity: | 97% | ||
| Prep. Method: | Gilbert, E. J., and Maxwell, A. (1994) Mol. Microbiol. 12, 365-373. | ||
| Name: | DNA gyrase subunit B | ||
| Synonyms: | gyrB | ||
| Type: | Enzyme | ||
| Mol. Mass.: | 89941.28 | ||
| Organism: | Escherichia coli | ||
| Description: | C3SLN3 | ||
| Residue: | 804 | ||
| Sequence: |
| ||
| BDBM283 | |||
| Source: | Synthesis | ||
| Purity: | n/a | ||
| Prep. Method: | Ferroud, D., Collard, J., Klich, M., Dupuis-Hamelin, C., Mauvais, P., Lassaigne, P., Bonnefoy, A., and Musicki, B.(1999) Bioorg. Med. Chem. Lett. 9, 2881-2886. | ||
| Name | BDBM283 | ||
| Synonyms: | 6-[(8-chloro-4-hydroxy-3-{[4-hydroxy-3-(3-methylbut-2-en-1-yl)benzene]amido}-2-oxo-2H-chromen-7-yl)oxy]-5-hydroxy-3-methoxy-2,2-dimethyloxan-4-yl 5-methyl-1H-pyrrole-2-carboxylate | Clorobiocin | Coumarin-Based, DNA Gyrase Inhibitor | cid_54677920 | ||
| Type | Antibiotic | ||
| Emp. Form. | C35H37ClN2O11 | ||
| Mol. Mass. | 697.128 | ||
| SMILES | COC1C(OC(=O)c2ccc(C)[nH]2)C(O)C(Oc2ccc3c(O)c(NC(=O)c4ccc(O)c(CC=C(C)C)c4)c(=O)oc3c2Cl)OC1(C)C |(-11.67,5.02,;-11.54,3.68,;-10.41,2.64,;-9.47,1.42,;-9.65,-.11,;-9.68,-1.17,;-8.71,-2.36,;-10.68,-1.93,;-11.03,-.43,;-13.62,-1.04,;-13.27,-2.54,;-14.39,-3.59,;-11.8,-2.98,;-7.94,1.61,;-8.07,.08,;-7.35,3.03,;-6.07,3.72,;-4.54,3.57,;-4.54,2.03,;-3.21,1.26,;-1.87,2.03,;-.54,1.26,;-.54,-.28,;.79,2.03,;1.98,1.05,;3.09,1.05,;3.55,-.42,;4.14,1.87,;4.14,3.41,;5.48,4.18,;6.81,3.41,;7.98,4.41,;6.81,1.87,;7.64,1.05,;8.4,1.75,;9.16,.93,;9.22,-.17,;10.21,1.4,;5.48,1.1,;.79,3.57,;2.05,4.46,;-.54,4.34,;-1.87,3.57,;-3.21,4.34,;-3.1,6.11,;-8.29,4.26,;-9.81,4.06,;-9.92,5.25,;-9.68,3.15,)| | ||
| Structure | 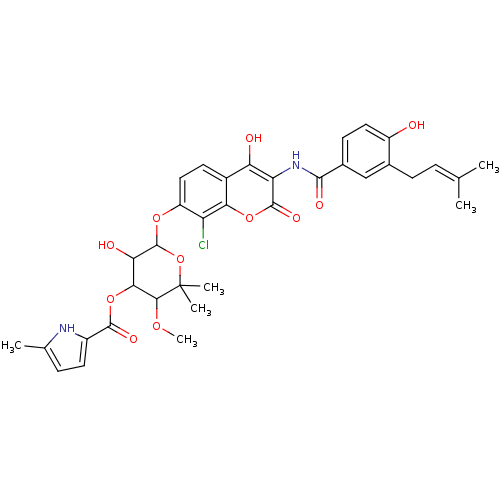
| ||
|
Home |
| |
Search |
| |
Deposit |
| |
SiteMap |
| |
About us |
| |
Email us |
| |
Info |
|
©2000 BindingDB. All rights reserved. |
|||||||||||||