| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Fibroblast growth factor receptor 2 |
|---|
| Ligand | BDBM339807 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | FGFR2 (Enzymatic Assay) |
|---|
| IC50 | 0.126±n/a nM |
|---|
| Citation |  Angibaud, PR; Obringer, M; Marin, JJ; Jeanty, M Naphthyridine derivative compounds US Patent US9757364 Publication Date 9/12/2017 Angibaud, PR; Obringer, M; Marin, JJ; Jeanty, M Naphthyridine derivative compounds US Patent US9757364 Publication Date 9/12/2017 |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Fibroblast growth factor receptor 2 |
|---|
| Name: | Fibroblast growth factor receptor 2 |
|---|
| Synonyms: | BEK | CD_antigen=CD332 | FGFR-2 | FGFR-2 Tyrosine Kinase | FGFR2 | FGFR2_HUMAN | Fibroblast growth factor receptor 2 (FGFR2) | Fibroblast growth factor receptor 2 precursor | KGFR | KSAM | Keratinocyte growth factor receptor | Keratinocyte growth factor receptor 2 | VEGF-receptor 2 and Fibroblast growth factor receptor 2 |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 92015.45 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P21802 |
|---|
| Residue: | 821 |
|---|
| Sequence: | MVSWGRFICLVVVTMATLSLARPSFSLVEDTTLEPEEPPTKYQISQPEVYVAAPGESLEV
RCLLKDAAVISWTKDGVHLGPNNRTVLIGEYLQIKGATPRDSGLYACTASRTVDSETWYF
MVNVTDAISSGDDEDDTDGAEDFVSENSNNKRAPYWTNTEKMEKRLHAVPAANTVKFRCP
AGGNPMPTMRWLKNGKEFKQEHRIGGYKVRNQHWSLIMESVVPSDKGNYTCVVENEYGSI
NHTYHLDVVERSPHRPILQAGLPANASTVVGGDVEFVCKVYSDAQPHIQWIKHVEKNGSK
YGPDGLPYLKVLKAAGVNTTDKEIEVLYIRNVTFEDAGEYTCLAGNSIGISFHSAWLTVL
PAPGREKEITASPDYLEIAIYCIGVFLIACMVVTVILCRMKNTTKKPDFSSQPAVHKLTK
RIPLRRQVTVSAESSSSMNSNTPLVRITTRLSSTADTPMLAGVSEYELPEDPKWEFPRDK
LTLGKPLGEGCFGQVVMAEAVGIDKDKPKEAVTVAVKMLKDDATEKDLSDLVSEMEMMKM
IGKHKNIINLLGACTQDGPLYVIVEYASKGNLREYLRARRPPGMEYSYDINRVPEEQMTF
KDLVSCTYQLARGMEYLASQKCIHRDLAARNVLVTENNVMKIADFGLARDINNIDYYKKT
TNGRLPVKWMAPEALFDRVYTHQSDVWSFGVLMWEIFTLGGSPYPGIPVEELFKLLKEGH
RMDKPANCTNELYMMMRDCWHAVPSQRPTFKQLVEDLDRILTLTTNEEYLDLSQPLEQYS
PSYPDTRSSCSSGDDSVFSPDPMPYEPCLPQYPHINGSVKT
|
|
|
|---|
| BDBM339807 |
|---|
| n/a |
|---|
| Name | BDBM339807 |
|---|
| Synonyms: | US9757364, Compound 11 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C23H24ClN5O3 |
|---|
| Mol. Mass. | 453.921 |
|---|
| SMILES | COc1cc(OC)cc(c1)N(CCCO)c1ccc2ncc(-c3cnn(C)c3)c(Cl)c2n1 |
|---|
| Structure | 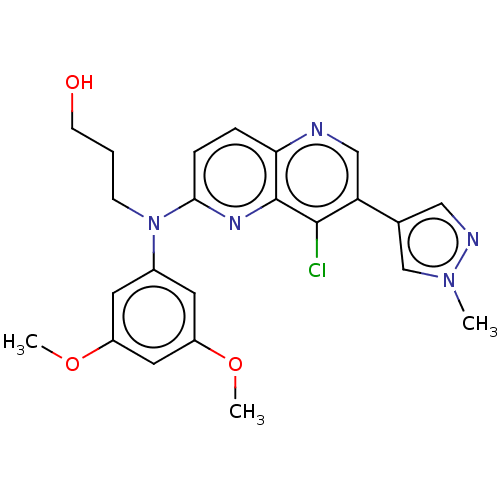
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Angibaud, PR; Obringer, M; Marin, JJ; Jeanty, M Naphthyridine derivative compounds US Patent US9757364 Publication Date 9/12/2017
Angibaud, PR; Obringer, M; Marin, JJ; Jeanty, M Naphthyridine derivative compounds US Patent US9757364 Publication Date 9/12/2017