

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | E3 ubiquitin-protein ligase Mdm2 [17-125] | ||
| Ligand | BDBM227651 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | FRET Assay | ||
| Temperature | 298.15±n/a K | ||
| IC50 | 0.176±n/a nM | ||
| Comments | extracted | ||
| Citation |  Cammarano, CM; Christopher, MP; Dinsmore, C; Doll, RJ; Fradera Llinas, FX; Li, C; Machacek, M; Martinez, M; Nair, LG; Pan, W; Reutershan, MH; Shizuka, M; Steinhuebel, DP; Sun, B; Thompson, CF; Trotter, BW; Wang, Y; Yang, L; Bogen, SL; Voss, ME; Panda, J; Kurissery, AT 2,6,7,8 substituted purines as HDM2 inhibitors US Patent US9540377 Publication Date 1/10/2017 Cammarano, CM; Christopher, MP; Dinsmore, C; Doll, RJ; Fradera Llinas, FX; Li, C; Machacek, M; Martinez, M; Nair, LG; Pan, W; Reutershan, MH; Shizuka, M; Steinhuebel, DP; Sun, B; Thompson, CF; Trotter, BW; Wang, Y; Yang, L; Bogen, SL; Voss, ME; Panda, J; Kurissery, AT 2,6,7,8 substituted purines as HDM2 inhibitors US Patent US9540377 Publication Date 1/10/2017 | ||
| More Info.: | Get all data from this article, Assay Method | ||
| E3 ubiquitin-protein ligase Mdm2 [17-125] | |||
| Name: | E3 ubiquitin-protein ligase Mdm2 [17-125] | ||
| Synonyms: | Double minute 2 protein | E3 ubiquitin-protein ligase Mdm2 | MDM2 | MDM2_HUMAN | p53-binding protein Mdm2 | ||
| Type: | Enzyme Catalytic Domain | ||
| Mol. Mass.: | 12522.24 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | Residue 17 to 125 | ||
| Residue: | 109 | ||
| Sequence: |
| ||
| BDBM227651 | |||
| n/a | |||
| Name | BDBM227651 | ||
| Synonyms: | 6-(5-chloropyridin-3-yl)- 8-[1-fluoro-1-(2- fluorophenyl)ethyl]-7- [(trans-4- methylcyclohexyl)methyl]- 7h-purine-2-carboxylic acid (enantiomer 1) | US9540377, 23.66 | US9540377, 23.67 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C27H26ClF2N5O2 | ||
| Mol. Mass. | 525.977 | ||
| SMILES | C[C@H]1CC[C@H](Cn2c(nc3nc(nc(-c4cncc(Cl)c4)c23)C(O)=O)C(C)(F)c2ccccc2F)CC1 |r,wU:4.4,wD:1.0,(6.57,-3.25,;5.09,-2.85,;4.69,-1.36,;3.2,-.96,;2.11,-2.05,;.62,-1.65,;.23,-.17,;1.13,1.08,;.23,2.33,;-1.24,1.85,;-2.57,2.62,;-3.91,1.85,;-3.91,.31,;-2.57,-.46,;-2.57,-2,;-1.24,-2.77,;-1.24,-4.31,;-2.57,-5.08,;-3.91,-4.31,;-5.24,-5.08,;-3.91,-2.77,;-1.24,.31,;-5.24,2.62,;-6.57,1.85,;-5.24,4.16,;2.8,1.08,;3.57,-.25,;4.34,1.08,;3.57,2.41,;5.11,2.41,;5.88,3.75,;5.11,5.08,;3.57,5.08,;2.8,3.75,;1.26,3.75,;2.51,-3.54,;4,-3.94,)| | ||
| Structure | 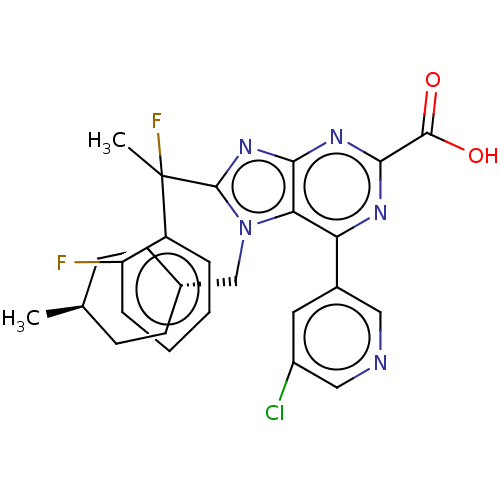
| ||