| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Purine nucleoside phosphorylase |
|---|
| Ligand | BDBM22103 |
|---|
| Substrate/Competitor | BDBM22104 |
|---|
| Meas. Tech. | PNP Inhibition Assay |
|---|
| pH | 7.7±n/a |
|---|
| Temperature | 295.15±n/a K |
|---|
| Ki | 0.665±0.06 nM |
|---|
| Km | 34000±n/a nM |
|---|
| Comments | Ki is the dissociation constant for the first step in the two-step binding characteristic of slow-onset tight-binding inhibition. |
|---|
| Citation |  Evans, GB; Furneaux, RH; Greatrex, B; Murkin, AS; Schramm, VL; Tyler, PC Azetidine based transition state analogue inhibitors of N-ribosyl hydrolases and phosphorylases. J Med Chem51:948-56 (2008) [PubMed] Article Evans, GB; Furneaux, RH; Greatrex, B; Murkin, AS; Schramm, VL; Tyler, PC Azetidine based transition state analogue inhibitors of N-ribosyl hydrolases and phosphorylases. J Med Chem51:948-56 (2008) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| Purine nucleoside phosphorylase |
|---|
| Name: | Purine nucleoside phosphorylase |
|---|
| Synonyms: | Inosine phosphorylase | NP | PNP | PNPH_BOVIN | Purine Nucleoside Phosphorylase (PNP) | Purine nucleoside phosphorylase |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 32035.18 |
|---|
| Organism: | Bos taurus (bovine) |
|---|
| Description: | n/a |
|---|
| Residue: | 289 |
|---|
| Sequence: | MANGYTYEDYQDTAKWLLSHTEQRPQVAVICGSGLGGLVNKLTQAQTFDYSEIPNFPEST
VPGHAGRLVFGILNGRACVMMQGRFHMYEGYPFWKVTFPVRVFRLLGVETLVVTNAAGGL
NPNFEVGDIMLIRDHINLPGFSGENPLRGPNEERFGVRFPAMSDAYDRDMRQKAHSTWKQ
MGEQRELQEGTYVMLGGPNFETVAECRLLRNLGADAVGMSTVPEVIVARHCGLRVFGFSL
ITNKVIMDYESQGKANHEEVLEAGKQAAQKLEQFVSLLMASIPVSGHTG
|
|
|
|---|
| BDBM22103 |
|---|
| BDBM22104 |
|---|
| Name | BDBM22103 |
|---|
| Synonyms: | 7-{[3,3-bis(hydroxymethyl)azetidin-1-yl]methyl}-3H,4H,5H-pyrrolo[3,2-d]pyrimidin-4-one | Azetidine based compound, 34 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C12H16N4O3 |
|---|
| Mol. Mass. | 264.2804 |
|---|
| SMILES | OCC1(CO)CN(Cc2c[nH]c3c2nc[nH]c3=O)C1 |
|---|
| Structure | 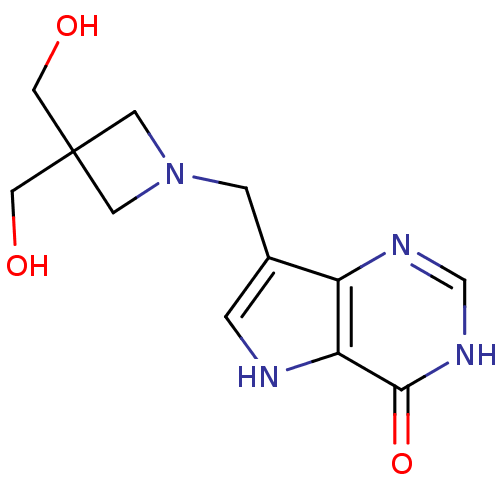
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Evans, GB; Furneaux, RH; Greatrex, B; Murkin, AS; Schramm, VL; Tyler, PC Azetidine based transition state analogue inhibitors of N-ribosyl hydrolases and phosphorylases. J Med Chem51:948-56 (2008) [PubMed] Article
Evans, GB; Furneaux, RH; Greatrex, B; Murkin, AS; Schramm, VL; Tyler, PC Azetidine based transition state analogue inhibitors of N-ribosyl hydrolases and phosphorylases. J Med Chem51:948-56 (2008) [PubMed] Article