| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Histamine H4 receptor |
|---|
| Ligand | BDBM22871 |
|---|
| Substrate/Competitor | BDBM7966 |
|---|
| Meas. Tech. | H4R Radioligand Binding Assay |
|---|
| pH | 7.4±n/a |
|---|
| Temperature | 310.15±n/a K |
|---|
| Ki | 3981±819 nM |
|---|
| Comments | Analysis of the [3H]histamine saturation binding yielded a Kd value of 11 +/- 1 and a Bmax value of 1.8 +/- 0.4 pmol/mg protein. |
|---|
| Citation |  Lim, HD; van Rijn, RM; Ling, P; Bakker, RA; Thurmond, RL; Leurs, R Evaluation of histamine H1-, H2-, and H3-receptor ligands at the human histamine H4 receptor: identification of 4-methylhistamine as the first potent and selective H4 receptor agonist. J Pharmacol Exp Ther314:1310-21 (2005) [PubMed] Article Lim, HD; van Rijn, RM; Ling, P; Bakker, RA; Thurmond, RL; Leurs, R Evaluation of histamine H1-, H2-, and H3-receptor ligands at the human histamine H4 receptor: identification of 4-methylhistamine as the first potent and selective H4 receptor agonist. J Pharmacol Exp Ther314:1310-21 (2005) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Inhibition_Run data, Solution Info, Assay Method |
|---|
| |
| Histamine H4 receptor |
|---|
| Name: | Histamine H4 receptor |
|---|
| Synonyms: | AXOR35 | G-protein coupled receptor 105 | GPCR105 | GPRv53 | HH4R | HISTAMINE H4 | HRH4 | HRH4_HUMAN | Histamine H4 receptor | Histamine H4 receptor (H4R) | Histamine receptor (H3 and H4) | Pfi-013 | SP9144 |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 44517.02 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Binding assays were using CHO cells stably expressing hH4R receptors. |
|---|
| Residue: | 390 |
|---|
| Sequence: | MPDTNSTINLSLSTRVTLAFFMSLVAFAIMLGNALVILAFVVDKNLRHRSSYFFLNLAIS
DFFVGVISIPLYIPHTLFEWDFGKEICVFWLTTDYLLCTASVYNIVLISYDRYLSVSNAV
SYRTQHTGVLKIVTLMVAVWVLAFLVNGPMILVSESWKDEGSECEPGFFSEWYILAITSF
LEFVIPVILVAYFNMNIYWSLWKRDHLSRCQSHPGLTAVSSNICGHSFRGRLSSRRSLSA
STEVPASFHSERQRRKSSLMFSSRTKMNSNTIASKMGSFSQSDSVALHQREHVELLRARR
LAKSLAILLGVFAVCWAPYSLFTIVLSFYSSATGPKSVWYRIAFWLQWFNSFVNPLLYPL
CHKRFQKAFLKIFCIKKQPLPSQHSRSVSS
|
|
|
|---|
| BDBM22871 |
|---|
| BDBM7966 |
|---|
| Name | BDBM22871 |
|---|
| Synonyms: | 13-chloro-10-(4-methylpiperazin-1-yl)-2-oxa-9-azatricyclo[9.4.0.0^{3,8}]pentadeca-1(11),3,5,7,9,12,14-heptaene | 2-chloro-11-(4-methylpiperazin-1-yl)dibenzo[b,f][1,4]oxazepine | 8-chloro-6-(4-methylpiperazino)benzo[b][1,4]benzoxazepine;succinic acid | CHEMBL831 | CL71,563 | Cloxazepine | Loxapine |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C18H18ClN3O |
|---|
| Mol. Mass. | 327.808 |
|---|
| SMILES | CN1CCN(CC1)C1=Nc2ccccc2Oc2ccc(Cl)cc12 |t:8| |
|---|
| Structure | 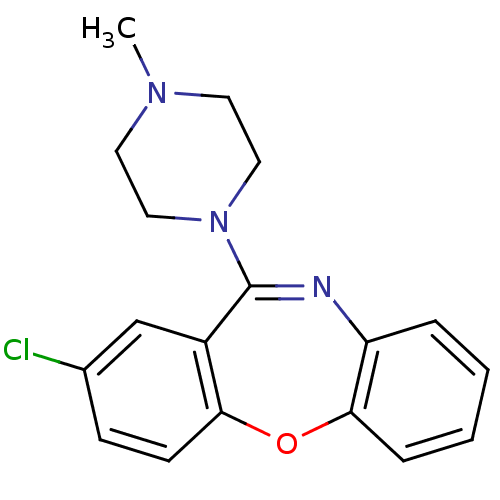
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Lim, HD; van Rijn, RM; Ling, P; Bakker, RA; Thurmond, RL; Leurs, R Evaluation of histamine H1-, H2-, and H3-receptor ligands at the human histamine H4 receptor: identification of 4-methylhistamine as the first potent and selective H4 receptor agonist. J Pharmacol Exp Ther314:1310-21 (2005) [PubMed] Article
Lim, HD; van Rijn, RM; Ling, P; Bakker, RA; Thurmond, RL; Leurs, R Evaluation of histamine H1-, H2-, and H3-receptor ligands at the human histamine H4 receptor: identification of 4-methylhistamine as the first potent and selective H4 receptor agonist. J Pharmacol Exp Ther314:1310-21 (2005) [PubMed] Article