| Reaction Details |
|---|
 | Report a problem with these data |
| Target | DNA ligase 1 |
|---|
| Ligand | BDBM23044 |
|---|
| Substrate/Competitor | BDBM23028 |
|---|
| Meas. Tech. | FAAH Inhibition Assay |
|---|
| pH | 9±n/a |
|---|
| Temperature | 295.15±n/a K |
|---|
| Ki | 5.6±0.2 nM |
|---|
| Citation |  Hardouin, C; Kelso, MJ; Romero, FA; Rayl, TJ; Leung, D; Hwang, I; Cravatt, BF; Boger, DL Structure-activity relationships of alpha-ketooxazole inhibitors of fatty acid amide hydrolase. J Med Chem50:3359-68 (2007) [PubMed] Article Hardouin, C; Kelso, MJ; Romero, FA; Rayl, TJ; Leung, D; Hwang, I; Cravatt, BF; Boger, DL Structure-activity relationships of alpha-ketooxazole inhibitors of fatty acid amide hydrolase. J Med Chem50:3359-68 (2007) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Inhibition_Run data, Solution Info, Assay Method |
|---|
| |
| DNA ligase 1 |
|---|
| Name: | DNA ligase 1 |
|---|
| Synonyms: | ATP-dependent DNA ligase | DNA Ligase I | DNLI1_HUMAN | Fatty Acid Amide Hydrolase | LIG1 | Polydeoxyribonucleotide synthase [ATP] |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 101721.76 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | n/a |
|---|
| Residue: | 919 |
|---|
| Sequence: | MQRSIMSFFHPKKEGKAKKPEKEASNSSRETEPPPKAALKEWNGVVSESDSPVKRPGRKA
ARVLGSEGEEEDEALSPAKGQKPALDCSQVSPPRPATSPENNASLSDTSPMDSSPSGIPK
RRTARKQLPKRTIQEVLEEQSEDEDREAKRKKEEEEEETPKESLTEAEVATEKEGEDGDQ
PTTPPKPLKTSKAETPTESVSEPEVATKQELQEEEEQTKPPRRAPKTLSSFFTPRKPAVK
KEVKEEEPGAPGKEGAAEGPLDPSGYNPAKNNYHPVEDACWKPGQKVPYLAVARTFEKIE
EVSARLRMVETLSNLLRSVVALSPPDLLPVLYLSLNHLGPPQQGLELGVGDGVLLKAVAQ
ATGRQLESVRAEAAEKGDVGLVAENSRSTQRLMLPPPPLTASGVFSKFRDIARLTGSAST
AKKIDIIKGLFVACRHSEARFIARSLSGRLRLGLAEQSVLAALSQAVSLTPPGQEFPPAM
VDAGKGKTAEARKTWLEEQGMILKQTFCEVPDLDRIIPVLLEHGLERLPEHCKLSPGIPL
KPMLAHPTRGISEVLKRFEEAAFTCEYKYDGQRAQIHALEGGEVKIFSRNQEDNTGKYPD
IISRIPKIKLPSVTSFILDTEAVAWDREKKQIQPFQVLTTRKRKEVDASEIQVQVCLYAF
DLIYLNGESLVREPLSRRRQLLRENFVETEGEFVFATSLDTKDIEQIAEFLEQSVKDSCE
GLMVKTLDVDATYEIAKRSHNWLKLKKDYLDGVGDTLDLVVIGAYLGRGKRAGRYGGFLL
ASYDEDSEELQAICKLGTGFSDEELEEHHQSLKALVLPSPRPYVRIDGAVIPDHWLDPSA
VWEVKCADLSLSPIYPAARGLVDSDKGISLRFPRFIRVREDKQPEQATTSAQVACLYRKQ
SQIQNQQGEDSGSDPEDTY
|
|
|
|---|
| BDBM23044 |
|---|
| BDBM23028 |
|---|
| Name | BDBM23044 |
|---|
| Synonyms: | alpha-ketooxazole, 5p | tert-butyl N-(4-{7-oxo-7-[5-(pyridin-2-yl)-1,3-oxazol-2-yl]heptyl}phenyl)carbamate |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C26H31N3O4 |
|---|
| Mol. Mass. | 449.542 |
|---|
| SMILES | CC(C)(C)OC(=O)Nc1ccc(CCCCCCC(=O)c2ncc(o2)-c2ccccn2)cc1 |
|---|
| Structure | 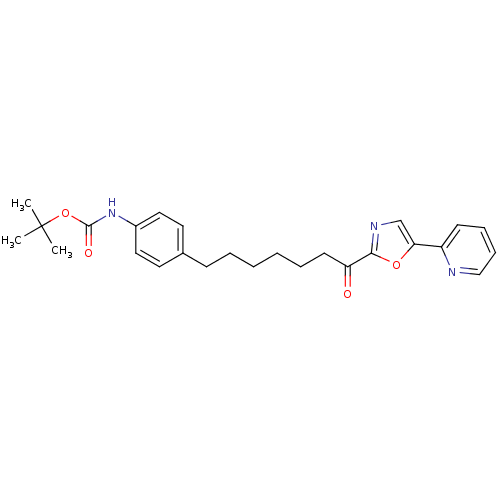
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Hardouin, C; Kelso, MJ; Romero, FA; Rayl, TJ; Leung, D; Hwang, I; Cravatt, BF; Boger, DL Structure-activity relationships of alpha-ketooxazole inhibitors of fatty acid amide hydrolase. J Med Chem50:3359-68 (2007) [PubMed] Article
Hardouin, C; Kelso, MJ; Romero, FA; Rayl, TJ; Leung, D; Hwang, I; Cravatt, BF; Boger, DL Structure-activity relationships of alpha-ketooxazole inhibitors of fatty acid amide hydrolase. J Med Chem50:3359-68 (2007) [PubMed] Article