| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Poly [ADP-ribose] polymerase 1 |
|---|
| Ligand | BDBM27695 |
|---|
| Substrate/Competitor | BDBM27683 |
|---|
| Meas. Tech. | PARP-1 Enzyme Inhibition Assay |
|---|
| pH | 8±n/a |
|---|
| Temperature | 296.15±n/a K |
|---|
| IC50 | 106±n/a nM |
|---|
| Citation |  Moree, WJ; Goldman, P; Demaggio, AJ; Christenson, E; Herendeen, D; Eksterowicz, J; Kesicki, EA; McElligott, DL; Beaton, G Identification of ring-fused pyrazolo pyridin-2-ones as novel poly(ADP-ribose)polymerase-1 inhibitors. Bioorg Med Chem Lett18:5126-9 (2008) [PubMed] Article Moree, WJ; Goldman, P; Demaggio, AJ; Christenson, E; Herendeen, D; Eksterowicz, J; Kesicki, EA; McElligott, DL; Beaton, G Identification of ring-fused pyrazolo pyridin-2-ones as novel poly(ADP-ribose)polymerase-1 inhibitors. Bioorg Med Chem Lett18:5126-9 (2008) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| Poly [ADP-ribose] polymerase 1 |
|---|
| Name: | Poly [ADP-ribose] polymerase 1 |
|---|
| Synonyms: | (ARTD1 or PARP1) | 2.4.2.- | 2.4.2.30 | ADP-ribosyltransferase diphtheria toxin-like 1 | ADPRT | ADPRT 1 | ARTD1 | DNA ADP-ribosyltransferase PARP1 | Human diphtheria toxin-like ADP-ribosyltransferase (ARTD1 or PARP1) | NAD(+) ADP-ribosyltransferase 1 | NT-PARP-1 | PARP-1 | PARP1 | PARP1_HUMAN | PPOL | Poly [ADP-ribose] polymerase (PARP) | Poly [ADP-ribose] polymerase 1 (PARP) | Poly [ADP-ribose] polymerase 1 (PARP-1) | Poly [ADP-ribose] polymerase 1 (PARP1) | Poly [ADP-ribose] polymerase 1, 24-kDa form | Poly [ADP-ribose] polymerase 1, 28-kDa form | Poly [ADP-ribose] polymerase 1, 89-kDa form | Poly [ADP-ribose] polymerase 1, processed C-terminus | Poly [ADP-ribose] polymerase 1, processed N-terminus | Poly [ADP-ribose] polymerase-1 | Poly(ADP-ribose) polymerase 1 (PARP1) | Poly(ADP-ribose) polymerase-1 (ARTD1/PARP1) | Poly[ADP-ribose] synthase 1 | Protein poly-ADP-ribosyltransferase PARP1 | Synonyms=ADPRT |
|---|
| Type: | n/a |
|---|
| Mol. Mass.: | 113114.22 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P09874 |
|---|
| Residue: | 1014 |
|---|
| Sequence: | MAESSDKLYRVEYAKSGRASCKKCSESIPKDSLRMAIMVQSPMFDGKVPHWYHFSCFWKV
GHSIRHPDVEVDGFSELRWDDQQKVKKTAEAGGVTGKGQDGIGSKAEKTLGDFAAEYAKS
NRSTCKGCMEKIEKGQVRLSKKMVDPEKPQLGMIDRWYHPGCFVKNREELGFRPEYSASQ
LKGFSLLATEDKEALKKQLPGVKSEGKRKGDEVDGVDEVAKKKSKKEKDKDSKLEKALKA
QNDLIWNIKDELKKVCSTNDLKELLIFNKQQVPSGESAILDRVADGMVFGALLPCEECSG
QLVFKSDAYYCTGDVTAWTKCMVKTQTPNRKEWVTPKEFREISYLKKLKVKKQDRIFPPE
TSASVAATPPPSTASAPAAVNSSASADKPLSNMKILTLGKLSRNKDEVKAMIEKLGGKLT
GTANKASLCISTKKEVEKMNKKMEEVKEANIRVVSEDFLQDVSASTKSLQELFLAHILSP
WGAEVKAEPVEVVAPRGKSGAALSKKSKGQVKEEGINKSEKRMKLTLKGGAAVDPDSGLE
HSAHVLEKGGKVFSATLGLVDIVKGTNSYYKLQLLEDDKENRYWIFRSWGRVGTVIGSNK
LEQMPSKEDAIEHFMKLYEEKTGNAWHSKNFTKYPKKFYPLEIDYGQDEEAVKKLTVNPG
TKSKLPKPVQDLIKMIFDVESMKKAMVEYEIDLQKMPLGKLSKRQIQAAYSILSEVQQAV
SQGSSDSQILDLSNRFYTLIPHDFGMKKPPLLNNADSVQAKVEMLDNLLDIEVAYSLLRG
GSDDSSKDPIDVNYEKLKTDIKVVDRDSEEAEIIRKYVKNTHATTHNAYDLEVIDIFKIE
REGECQRYKPFKQLHNRRLLWHGSRTTNFAGILSQGLRIAPPEAPVTGYMFGKGIYFADM
VSKSANYCHTSQGDPIGLILLGEVALGNMYELKHASHISKLPKGKHSVKGLGKTTPDPSA
NISLDGVDVPLGTGISSGVNDTSLLYNEYIVYDIAQVNLKYLLKLKFNFKTSLW
|
|
|
|---|
| BDBM27695 |
|---|
| BDBM27683 |
|---|
| Name | BDBM27695 |
|---|
| Synonyms: | 5-methyl-3-phenyl-4,5,7-triazatricyclo[7.3.0.0^{2,6}]dodeca-1(9),2(6),3-trien-8-one | ring-fused pyrazolo pyridin-2-one, 22 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C16H15N3O |
|---|
| Mol. Mass. | 265.3098 |
|---|
| SMILES | Cn1nc(-c2ccccc2)c2c3CCCc3c(=O)[nH]c12 |
|---|
| Structure | 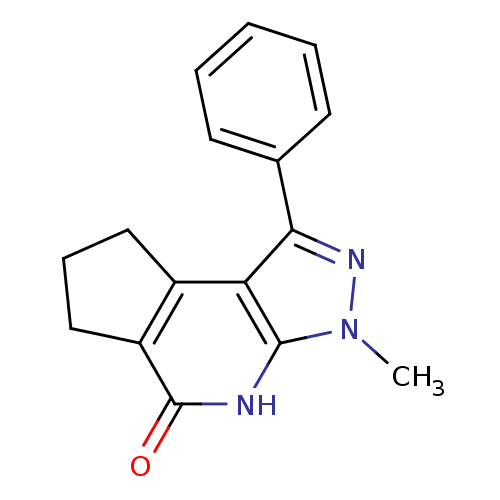
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Moree, WJ; Goldman, P; Demaggio, AJ; Christenson, E; Herendeen, D; Eksterowicz, J; Kesicki, EA; McElligott, DL; Beaton, G Identification of ring-fused pyrazolo pyridin-2-ones as novel poly(ADP-ribose)polymerase-1 inhibitors. Bioorg Med Chem Lett18:5126-9 (2008) [PubMed] Article
Moree, WJ; Goldman, P; Demaggio, AJ; Christenson, E; Herendeen, D; Eksterowicz, J; Kesicki, EA; McElligott, DL; Beaton, G Identification of ring-fused pyrazolo pyridin-2-ones as novel poly(ADP-ribose)polymerase-1 inhibitors. Bioorg Med Chem Lett18:5126-9 (2008) [PubMed] Article