| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Cannabinoid receptor 1 |
|---|
| Ligand | BDBM29061 |
|---|
| Substrate/Competitor | BDBM21278 |
|---|
| Meas. Tech. | CB1 Radioligand Binding Assay (Ki) and GTP-gamma-[35S] Binding Assay (EC50) |
|---|
| pH | 7.4±n/a |
|---|
| Temperature | 303.15±n/a K |
|---|
| Ki | 0.6±0.1 nM |
|---|
| EC50 | 1.0±n/a nM |
|---|
| Citation |  Dow, RL; Carpino, PA; Hadcock, JR; Black, SC; Iredale, PA; DaSilva-Jardine, P; Schneider, SR; Paight, ES; Griffith, DA; Scott, DO; O'Connor, RE; Nduaka, CI Discovery of 2-(2-chlorophenyl)-3-(4-chlorophenyl)-7-(2,2-difluoropropyl)-6,7-dihydro-2H-pyrazolo[3,4-f][1,4]oxazepin-8(5H)-one (PF-514273), a novel, bicyclic lactam-based cannabinoid-1 receptor antagonist for the treatment of obesity. J Med Chem52:2652-5 (2009) [PubMed] Article Dow, RL; Carpino, PA; Hadcock, JR; Black, SC; Iredale, PA; DaSilva-Jardine, P; Schneider, SR; Paight, ES; Griffith, DA; Scott, DO; O'Connor, RE; Nduaka, CI Discovery of 2-(2-chlorophenyl)-3-(4-chlorophenyl)-7-(2,2-difluoropropyl)-6,7-dihydro-2H-pyrazolo[3,4-f][1,4]oxazepin-8(5H)-one (PF-514273), a novel, bicyclic lactam-based cannabinoid-1 receptor antagonist for the treatment of obesity. J Med Chem52:2652-5 (2009) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| Cannabinoid receptor 1 |
|---|
| Name: | Cannabinoid receptor 1 |
|---|
| Synonyms: | CANN6 | CANNABINOID CB1 | CB-R | CB1 | CNR | CNR1 | CNR1_HUMAN | Cannabinoid CB1 receptor | Cannabinoid receptor | Cannabinoid receptor 1 (CB1) | Cannabinoid receptor 1 (brain) |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 52868.96 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P21554 |
|---|
| Residue: | 472 |
|---|
| Sequence: | MKSILDGLADTTFRTITTDLLYVGSNDIQYEDIKGDMASKLGYFPQKFPLTSFRGSPFQE
KMTAGDNPQLVPADQVNITEFYNKSLSSFKENEENIQCGENFMDIECFMVLNPSQQLAIA
VLSLTLGTFTVLENLLVLCVILHSRSLRCRPSYHFIGSLAVADLLGSVIFVYSFIDFHVF
HRKDSRNVFLFKLGGVTASFTASVGSLFLTAIDRYISIHRPLAYKRIVTRPKAVVAFCLM
WTIAIVIAVLPLLGWNCEKLQSVCSDIFPHIDETYLMFWIGVTSVLLLFIVYAYMYILWK
AHSHAVRMIQRGTQKSIIIHTSEDGKVQVTRPDQARMDIRLAKTLVLILVVLIICWGPLL
AIMVYDVFGKMNKLIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMFPSCEGTAQ
PLDNSMGDSDCLHKHANNAASVHRAAESCIKSTVKIAKVTMSVSTDTSAEAL
|
|
|
|---|
| BDBM29061 |
|---|
| BDBM21278 |
|---|
| Name | BDBM29061 |
|---|
| Synonyms: | CHEMBL201602 | pyrazolopyrimidinone-based antagonist, 3 |
|---|
| Type | n/a |
|---|
| Emp. Form. | C20H13Cl2F3N4O |
|---|
| Mol. Mass. | 453.245 |
|---|
| SMILES | Cc1nc2c(-c3ccc(Cl)cc3)n(nc2c(=O)n1CC(F)(F)F)-c1ccccc1Cl |
|---|
| Structure | 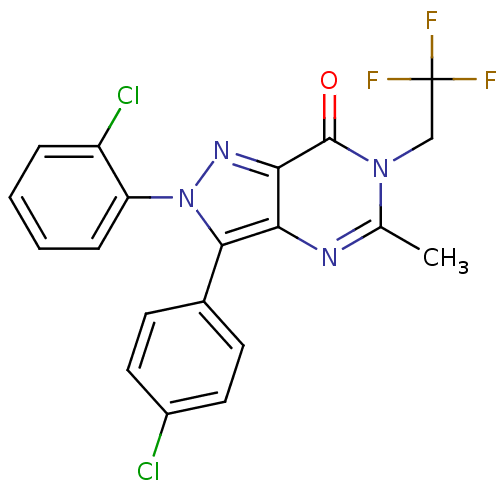
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Dow, RL; Carpino, PA; Hadcock, JR; Black, SC; Iredale, PA; DaSilva-Jardine, P; Schneider, SR; Paight, ES; Griffith, DA; Scott, DO; O'Connor, RE; Nduaka, CI Discovery of 2-(2-chlorophenyl)-3-(4-chlorophenyl)-7-(2,2-difluoropropyl)-6,7-dihydro-2H-pyrazolo[3,4-f][1,4]oxazepin-8(5H)-one (PF-514273), a novel, bicyclic lactam-based cannabinoid-1 receptor antagonist for the treatment of obesity. J Med Chem52:2652-5 (2009) [PubMed] Article
Dow, RL; Carpino, PA; Hadcock, JR; Black, SC; Iredale, PA; DaSilva-Jardine, P; Schneider, SR; Paight, ES; Griffith, DA; Scott, DO; O'Connor, RE; Nduaka, CI Discovery of 2-(2-chlorophenyl)-3-(4-chlorophenyl)-7-(2,2-difluoropropyl)-6,7-dihydro-2H-pyrazolo[3,4-f][1,4]oxazepin-8(5H)-one (PF-514273), a novel, bicyclic lactam-based cannabinoid-1 receptor antagonist for the treatment of obesity. J Med Chem52:2652-5 (2009) [PubMed] Article