| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Oxysterols receptor LXR-beta [154-461] |
|---|
| Ligand | BDBM20000 |
|---|
| Substrate/Competitor | BDBM19993 |
|---|
| Meas. Tech. | LXRbeta Binding Assay (IC50) and LAFbeta Functional Assay (EC50) |
|---|
| pH | 7.4±n/a |
|---|
| Temperature | 277.15±n/a K |
|---|
| IC50 | 2.1±n/a nM |
|---|
| EC50 | 71±n/a nM |
|---|
| Comments | Efficacy=97%. |
|---|
| Citation |  Bernotas, RC; Kaufman, DH; Singhaus, RR; Ullrich, J; Unwalla, R; Quinet, E; Nambi, P; Wilhelmsson, A; Goos-Nilsson, A; Wrobel, J 4-(3-aryloxyaryl)quinoline alcohols are liver X receptor agonists. Bioorg Med Chem17:8086-92 (2009) [PubMed] Article Bernotas, RC; Kaufman, DH; Singhaus, RR; Ullrich, J; Unwalla, R; Quinet, E; Nambi, P; Wilhelmsson, A; Goos-Nilsson, A; Wrobel, J 4-(3-aryloxyaryl)quinoline alcohols are liver X receptor agonists. Bioorg Med Chem17:8086-92 (2009) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| Oxysterols receptor LXR-beta [154-461] |
|---|
| Name: | Oxysterols receptor LXR-beta [154-461] |
|---|
| Synonyms: | LXRB | Liver X Receptor beta (LXR-beta) | NER | NR1H2 | NR1H2_HUMAN | Nuclear orphan receptor LXR-beta | Nuclear receptor NER | Nuclear receptor subfamily 1 group H member 2 | Oxysterols receptor LXR-beta | UNR | Ubiquitously-expressed nuclear receptor |
|---|
| Type: | Receptor |
|---|
| Mol. Mass.: | 34695.52 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | LXR beta ligand binding domain (amino acid residues 154-461) with an N-terminal biotinylation tag expressed in E.coli, was used for the binding assays. |
|---|
| Residue: | 308 |
|---|
| Sequence: | MREQCVLSEEQIRKKKIRKQQQQQESQSQSQSPVGPQGSSSSASGPGASPGGSEAGSQGS
GEGEGVQLTAAQELMIQQLVAAQLQCNKRSFSDQPKVTPWPLGADPQSRDARQQRFAHFT
ELAIISVQEIVDFAKQVPGFLQLGREDQIALLKASTIEIMLLETARRYNHETECITFLKD
FTYSKDDFHRAGLQVEFINPIFEFSRAMRRLGLDDAEYALLIAINIFSADRPNVQEPGRV
EALQQPYVEALLSYTRIKRPQDQLRFPRMLMKLVSLRTLSSVHSEQVFALRLQDKKLPPL
LSEIWDVH
|
|
|
|---|
| BDBM20000 |
|---|
| BDBM19993 |
|---|
| Name | BDBM20000 |
|---|
| Synonyms: | 2-(4-{3-[3-benzyl-8-(trifluoromethyl)quinolin-4-yl]phenoxymethyl}phenyl)acetic acid | CHEMBL214719 | JMC496151 Compound 15 | WAY-254011 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C32H24F3NO3 |
|---|
| Mol. Mass. | 527.5331 |
|---|
| SMILES | OC(=O)Cc1ccc(COc2cccc(c2)-c2c(Cc3ccccc3)cnc3c(cccc23)C(F)(F)F)cc1 |
|---|
| Structure | 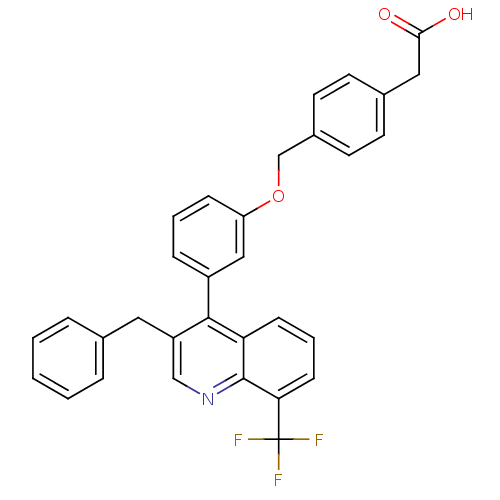
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Bernotas, RC; Kaufman, DH; Singhaus, RR; Ullrich, J; Unwalla, R; Quinet, E; Nambi, P; Wilhelmsson, A; Goos-Nilsson, A; Wrobel, J 4-(3-aryloxyaryl)quinoline alcohols are liver X receptor agonists. Bioorg Med Chem17:8086-92 (2009) [PubMed] Article
Bernotas, RC; Kaufman, DH; Singhaus, RR; Ullrich, J; Unwalla, R; Quinet, E; Nambi, P; Wilhelmsson, A; Goos-Nilsson, A; Wrobel, J 4-(3-aryloxyaryl)quinoline alcohols are liver X receptor agonists. Bioorg Med Chem17:8086-92 (2009) [PubMed] Article