| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Induced myeloid leukemia cell differentiation protein Mcl-1 |
|---|
| Ligand | BDBM78434 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Dose Response Confirmation for Mcl-1/Noxa Interaction Inhibitors |
|---|
| IC50 | 3773.276395±n/a nM |
|---|
| Citation |  PubChem, PC Dose Response Confirmation for Mcl-1/Noxa Interaction Inhibitors PubChem Bioassay(2008)[AID] PubChem, PC Dose Response Confirmation for Mcl-1/Noxa Interaction Inhibitors PubChem Bioassay(2008)[AID] |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| Induced myeloid leukemia cell differentiation protein Mcl-1 |
|---|
| Name: | Induced myeloid leukemia cell differentiation protein Mcl-1 |
|---|
| Synonyms: | BCL2L3 | Bcl-2-like protein 3 | Bcl-2-like protein 3 (Mcl-1) | Bcl-2-related protein EAT/mcl1 | Bcl2-L-3 | Induced myeloid leukemia cell differentiation protein (Mcl-1) | MCL1 | MCL1_HUMAN | Mcl-1 | Myeloid Cell factor-1 (Mcl-1) | Myeloid cell leukemia sequence 1 (BCL2-related) | mcl1/EAT |
|---|
| Type: | Membrane; Single-pass membrane protein |
|---|
| Mol. Mass.: | 37332.87 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q07820 |
|---|
| Residue: | 350 |
|---|
| Sequence: | MFGLKRNAVIGLNLYCGGAGLGAGSGGATRPGGRLLATEKEASARREIGGGEAGAVIGGS
AGASPPSTLTPDSRRVARPPPIGAEVPDVTATPARLLFFAPTRRAAPLEEMEAPAADAIM
SPEEELDGYEPEPLGKRPAVLPLLELVGESGNNTSTDGSLPSTPPPAEEEEDELYRQSLE
IISRYLREQATGAKDTKPMGRSGATSRKALETLRRVGDGVQRNHETAFQGMLRKLDIKNE
DDVKSLSRVMIHVFSDGVTNWGRIVTLISFGAFVAKHLKTINQESCIEPLAESITDVLVR
TKRDWLVKQRGWDGFVEFFHVEDLEGGIRNVLLAFAGVAGVGAGLAYLIR
|
|
|
|---|
| BDBM78434 |
|---|
| n/a |
|---|
| Name | BDBM78434 |
|---|
| Synonyms: | 2-chloranyl-10-[3-(4-methylpiperazin-1-yl)propyl]phenothiazine;ethane-1,2-disulfonic acid | 2-chloro-10-[3-(4-methyl-1-piperazinyl)propyl]phenothiazine;ethane-1,2-disulfonic acid | 2-chloro-10-[3-(4-methylpiperazin-1-yl)propyl]phenothiazine;ethane-1,2-disulfonic acid | 2-chloro-10-[3-(4-methylpiperazino)propyl]phenothiazine;ethane-1,2-disulfonic acid | Compazine | Compro | MLS001148133 | PROCHLORPERAZINE | SMR000653454 | cid_91499 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C20H24ClN3S |
|---|
| Mol. Mass. | 373.943 |
|---|
| SMILES | CN1CCN(CCCN2c3ccccc3Sc3ccc(Cl)cc23)CC1 |
|---|
| Structure | 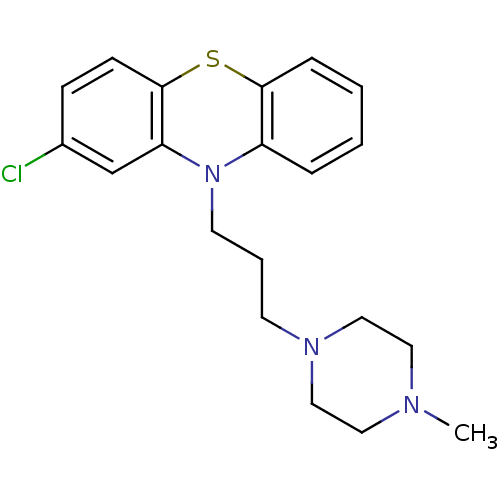
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 PubChem, PC Dose Response Confirmation for Mcl-1/Noxa Interaction Inhibitors PubChem Bioassay(2008)[AID]
PubChem, PC Dose Response Confirmation for Mcl-1/Noxa Interaction Inhibitors PubChem Bioassay(2008)[AID]