| Reaction Details |
|---|
 | Report a problem with these data |
| Target | ATP-dependent molecular chaperone HSP82 |
|---|
| Ligand | BDBM38924 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Fluorescence Cell-Based Secondary Assay to Identify Inhibitors of Resistant C. albicans Growth in the Presence of Fluconazole |
|---|
| EC50 | 521±1296 nM |
|---|
| Citation |  PubChem, PC Fluorescence Cell-Based Secondary Assay to Identify Inhibitors of Resistant C. albicans Growth in the Presence of Fluconazole PubChem Bioassay(2010)[AID] PubChem, PC Fluorescence Cell-Based Secondary Assay to Identify Inhibitors of Resistant C. albicans Growth in the Presence of Fluconazole PubChem Bioassay(2010)[AID] |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| ATP-dependent molecular chaperone HSP82 |
|---|
| Name: | ATP-dependent molecular chaperone HSP82 |
|---|
| Synonyms: | heat shock protein 90 |
|---|
| Type: | Enzyme Catalytic Domain |
|---|
| Mol. Mass.: | 80784.12 |
|---|
| Organism: | Candida albicans |
|---|
| Description: | gi_994798 |
|---|
| Residue: | 707 |
|---|
| Sequence: | MADAKVETHEFTAEISQLMSLIINTVYSNKEIFLRELISNASDALDKIRYQALSDPSQLE
SEPELFIRIIPQKDQKVLEIRDSGIGMTKADLVNNLGTIAKSGTKSFMEALSAGADVSMI
GQFGVGFYSLFLVADHVQVISKHNDDEQYVWESNAGGKFTVTLDETNERLGRGTMLRLFL
KEDQLEYLEEKRIKEVVKKHSEFVAYPIQLVVTKEVEKEVPETEEEDKAAEEDDKKPKLE
EVKDEEDEKKEKKTKTVKEEVTETEELNKTKPLWTRNPSDITQDEYNAFYKSISNDWEDP
LAVKHFSVEGQLEFRAILFVPKRAPFDAFESKKKKNNIKLYVRRVFITDDAEELIPEWLS
FIKGVVDSEDLPLNLSREMLQQNKILKVIRKNIVKKMIETFNEISEDQEQFNQFYTAFSK
NIKLGIHEDAQNRQSLAKLLRFYSTKSSEEMTSLSDYVTRMPEHQKNIYYITGESIKAVE
KSPFLDALKAKNFEVLFMVDPIDEYAMTQLKEFEDKKLVDITKDFELEESDEEKAAREKE
IKEYEPLTKALKDILGDQVEKVVVSYKLVDAPAAIRTGQFGWSANMERIMKAQALRDTTM
SSYMSSKKTFEISPSSPIIKELKKKVETDGAEDKTVKDLTTLLFDTALLTSGFTLDEPSN
FAHRINRLIALGLNIDDDSEETAVEPEATTTASTDEPAGESAMEEVD
|
|
|
|---|
| BDBM38924 |
|---|
| n/a |
|---|
| Name | BDBM38924 |
|---|
| Synonyms: | 2,6-bis(2-pyridinyl)pyridine | 2,6-bis(2-pyridyl)pyridine | 2,6-dipyridin-2-ylpyridine | MLS000048666 | SMR000060049 | cid_70848 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C15H11N3 |
|---|
| Mol. Mass. | 233.2679 |
|---|
| SMILES | c1ccc(nc1)-c1cccc(n1)-c1ccccn1 |
|---|
| Structure | 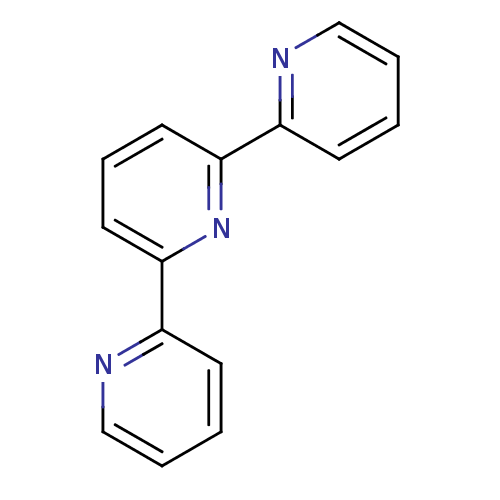
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 PubChem, PC Fluorescence Cell-Based Secondary Assay to Identify Inhibitors of Resistant C. albicans Growth in the Presence of Fluconazole PubChem Bioassay(2010)[AID]
PubChem, PC Fluorescence Cell-Based Secondary Assay to Identify Inhibitors of Resistant C. albicans Growth in the Presence of Fluconazole PubChem Bioassay(2010)[AID]