

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Interleukin-1 receptor-associated kinase 4 | ||
| Ligand | BDBM50246761 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_1676839 (CHEMBL4026982) | ||
| IC50 | 2.0±n/a nM | ||
| Citation |  Scott, JS; Degorce, SL; Anjum, R; Culshaw, J; Davies, RDM; Davies, NL; Dillman, KS; Dowling, JE; Drew, L; Ferguson, AD; Groombridge, SD; Halsall, CT; Hudson, JA; Lamont, S; Lindsay, NA; Marden, SK; Mayo, MF; Pease, JE; Perkins, DR; Pink, JH; Robb, GR; Rosen, A; Shen, M; McWhirter, C; Wu, D Discovery and Optimization of Pyrrolopyrimidine Inhibitors of Interleukin-1 Receptor Associated Kinase 4 (IRAK4) for the Treatment of Mutant MYD88 J Med Chem60:10071-10091 (2017) [PubMed] Article Scott, JS; Degorce, SL; Anjum, R; Culshaw, J; Davies, RDM; Davies, NL; Dillman, KS; Dowling, JE; Drew, L; Ferguson, AD; Groombridge, SD; Halsall, CT; Hudson, JA; Lamont, S; Lindsay, NA; Marden, SK; Mayo, MF; Pease, JE; Perkins, DR; Pink, JH; Robb, GR; Rosen, A; Shen, M; McWhirter, C; Wu, D Discovery and Optimization of Pyrrolopyrimidine Inhibitors of Interleukin-1 Receptor Associated Kinase 4 (IRAK4) for the Treatment of Mutant MYD88 J Med Chem60:10071-10091 (2017) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Interleukin-1 receptor-associated kinase 4 | |||
| Name: | Interleukin-1 receptor-associated kinase 4 | ||
| Synonyms: | IRAK-4 | IRAK4 | IRAK4_HUMAN | Interleukin-1 receptor-associated kinase 4 (IRAK-4) | Interleukin-1 receptor-associated kinase 4 (IRAK4) | Renal carcinoma antigen NY-REN-64 | ||
| Type: | Protein | ||
| Mol. Mass.: | 51519.08 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | Q9NWZ3 | ||
| Residue: | 460 | ||
| Sequence: |
| ||
| BDBM50246761 | |||
| n/a | |||
| Name | BDBM50246761 | ||
| Synonyms: | CHEMBL4102501 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C25H36N6O2 | ||
| Mol. Mass. | 452.5923 | ||
| SMILES | [H][C@@]12CC(=O)N(C)[C@]1([H])CCN(C2)[C@H]1CC[C@@H](CC1)Nc1ncnc2[nH]cc(C3CCOCC3)c12 |r,wU:16.21,1.0,7.8,wD:13.14,(14.1,-14.84,;15.43,-15.62,;15.76,-14.11,;17.29,-13.95,;18.06,-12.62,;17.91,-15.36,;19.41,-15.68,;16.76,-16.39,;18.09,-17.15,;16.75,-17.93,;15.42,-18.69,;14.1,-17.92,;14.09,-16.38,;12.76,-18.68,;11.43,-17.91,;10.09,-18.68,;10.1,-20.22,;11.43,-20.99,;12.75,-20.22,;8.76,-20.99,;8.77,-22.53,;10.1,-23.29,;10.11,-24.84,;8.77,-25.61,;7.44,-24.84,;5.97,-25.33,;5.05,-24.09,;5.95,-22.83,;5.47,-21.37,;6.48,-20.21,;6,-18.76,;4.5,-18.44,;3.47,-19.59,;3.95,-21.06,;7.43,-23.3,)| | ||
| Structure | 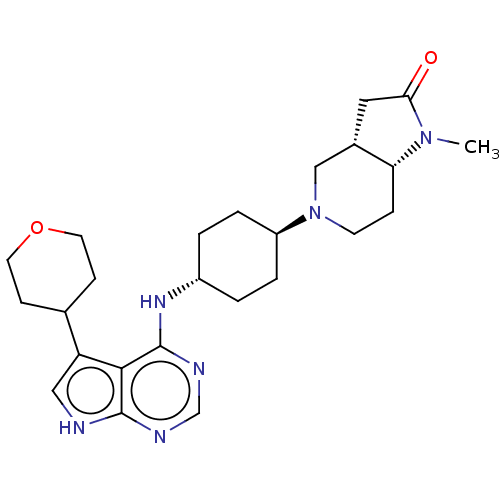
| ||