| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Beta-adrenergic receptor kinase 1 |
|---|
| Ligand | BDBM50257330 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1689808 (CHEMBL4040378) |
|---|
| EC50 | 12000±n/a nM |
|---|
| Citation |  Okawa, T; Aramaki, Y; Yamamoto, M; Kobayashi, T; Fukumoto, S; Toyoda, Y; Henta, T; Hata, A; Ikeda, S; Kaneko, M; Hoffman, ID; Sang, BC; Zou, H; Kawamoto, T Design, Synthesis, and Evaluation of the Highly Selective and Potent G-Protein-Coupled Receptor Kinase 2 (GRK2) Inhibitor for the Potential Treatment of Heart Failure. J Med Chem60:6942-6990 (2017) [PubMed] Article Okawa, T; Aramaki, Y; Yamamoto, M; Kobayashi, T; Fukumoto, S; Toyoda, Y; Henta, T; Hata, A; Ikeda, S; Kaneko, M; Hoffman, ID; Sang, BC; Zou, H; Kawamoto, T Design, Synthesis, and Evaluation of the Highly Selective and Potent G-Protein-Coupled Receptor Kinase 2 (GRK2) Inhibitor for the Potential Treatment of Heart Failure. J Med Chem60:6942-6990 (2017) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Beta-adrenergic receptor kinase 1 |
|---|
| Name: | Beta-adrenergic receptor kinase 1 |
|---|
| Synonyms: | ADRBK1 | ARBK1_HUMAN | BARK | BARK1 | Beta-ARK-1 | Beta-adrenergic receptor kinase 1 | G-protein coupled receptor kinase 2 | G-protein coupled receptor kinase 2 (GRK2) | GRK2 |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 79581.30 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P25098 |
|---|
| Residue: | 689 |
|---|
| Sequence: | MADLEAVLADVSYLMAMEKSKATPAARASKKILLPEPSIRSVMQKYLEDRGEVTFEKIFS
QKLGYLLFRDFCLNHLEEARPLVEFYEEIKKYEKLETEEERVARSREIFDSYIMKELLAC
SHPFSKSATEHVQGHLGKKQVPPDLFQPYIEEICQNLRGDVFQKFIESDKFTRFCQWKNV
ELNIHLTMNDFSVHRIIGRGGFGEVYGCRKADTGKMYAMKCLDKKRIKMKQGETLALNER
IMLSLVSTGDCPFIVCMSYAFHTPDKLSFILDLMNGGDLHYHLSQHGVFSEADMRFYAAE
IILGLEHMHNRFVVYRDLKPANILLDEHGHVRISDLGLACDFSKKKPHASVGTHGYMAPE
VLQKGVAYDSSADWFSLGCMLFKLLRGHSPFRQHKTKDKHEIDRMTLTMAVELPDSFSPE
LRSLLEGLLQRDVNRRLGCLGRGAQEVKESPFFRSLDWQMVFLQKYPPPLIPPRGEVNAA
DAFDIGSFDEEDTKGIKLLDSDQELYRNFPLTISERWQQEVAETVFDTINAETDRLEARK
KAKNKQLGHEEDYALGKDCIMHGYMSKMGNPFLTQWQRRYFYLFPNRLEWRGEGEAPQSL
LTMEEIQSVEETQIKERKCLLLKIRGGKQFILQCDSDPELVQWKKELRDAYREAQQLVQR
VPKMKNKPRSPVVELSKVPLVQRGSANGL
|
|
|
|---|
| BDBM50257330 |
|---|
| n/a |
|---|
| Name | BDBM50257330 |
|---|
| Synonyms: | CHEMBL4071398 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C25H23F3N6O2 |
|---|
| Mol. Mass. | 496.4843 |
|---|
| SMILES | OCCn1c(CNc2cccc(c2)C(=O)NCc2ccccc2C(F)(F)F)nnc1-c1ccncc1 |
|---|
| Structure | 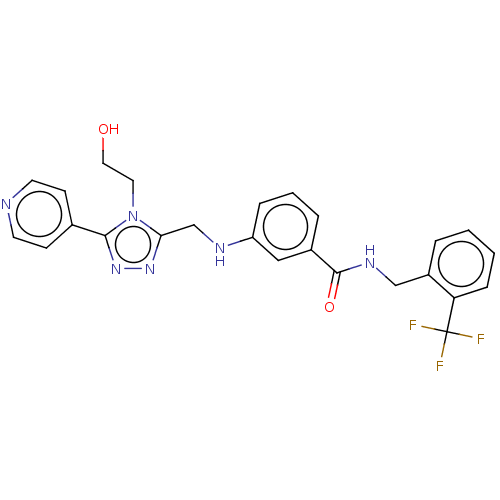
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Okawa, T; Aramaki, Y; Yamamoto, M; Kobayashi, T; Fukumoto, S; Toyoda, Y; Henta, T; Hata, A; Ikeda, S; Kaneko, M; Hoffman, ID; Sang, BC; Zou, H; Kawamoto, T Design, Synthesis, and Evaluation of the Highly Selective and Potent G-Protein-Coupled Receptor Kinase 2 (GRK2) Inhibitor for the Potential Treatment of Heart Failure. J Med Chem60:6942-6990 (2017) [PubMed] Article
Okawa, T; Aramaki, Y; Yamamoto, M; Kobayashi, T; Fukumoto, S; Toyoda, Y; Henta, T; Hata, A; Ikeda, S; Kaneko, M; Hoffman, ID; Sang, BC; Zou, H; Kawamoto, T Design, Synthesis, and Evaluation of the Highly Selective and Potent G-Protein-Coupled Receptor Kinase 2 (GRK2) Inhibitor for the Potential Treatment of Heart Failure. J Med Chem60:6942-6990 (2017) [PubMed] Article