| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Adenosine receptor A2a |
|---|
| Ligand | BDBM50004566 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_31063 (CHEMBL641347) |
|---|
| Ki | 1.2±n/a nM |
|---|
| Citation |  Baraldi, PG; Cacciari, B; Spalluto, G; Pineda de las Infantas y Villatoro, MJ; Zocchi, C; Dionisotti, S; Ongini, E Pyrazolo[4,3-e]-1,2,4-triazolo[1,5-c]pyrimidine derivatives: potent and selective A(2A) adenosine antagonists. J Med Chem39:1164-71 (1996) [PubMed] Article Baraldi, PG; Cacciari, B; Spalluto, G; Pineda de las Infantas y Villatoro, MJ; Zocchi, C; Dionisotti, S; Ongini, E Pyrazolo[4,3-e]-1,2,4-triazolo[1,5-c]pyrimidine derivatives: potent and selective A(2A) adenosine antagonists. J Med Chem39:1164-71 (1996) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Adenosine receptor A2a |
|---|
| Name: | Adenosine receptor A2a |
|---|
| Synonyms: | AA2AR_RAT | ADENOSINE A2a | Adenosine A2 receptor | Adenosine A2a receptor (A2a) | Adenosine Receptors A2a (A2a) | Adenosine receptor A2a and A3 | Adenosine receptors A2a | Adora2a | Rat striatal adenosine A2a receptor |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 45015.65 |
|---|
| Organism: | Rattus norvegicus (rat) |
|---|
| Description: | Rat A2A receptors expressed in CHO cells. |
|---|
| Residue: | 410 |
|---|
| Sequence: | MGSSVYITVELAIAVLAILGNVLVCWAVWINSNLQNVTNFFVVSLAAADIAVGVLAIPFA
ITISTGFCAACHGCLFFACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGVRAKG
IIAICWVLSFAIGLTPMLGWNNCSQKDGNSTKTCGEGRVTCLFEDVVPMNYMVYYNFFAF
VLLPLLLMLAIYLRIFLAARRQLKQMESQPLPGERTRSTLQKEVHAAKSLAIIVGLFALC
WLPLHIINCFTFFCSTCRHAPPWLMYLAIILSHSNSVVNPFIYAYRIREFRQTFRKIIRT
HVLRRQEPFQAGGSSAWALAAHSTEGEQVSLRLNGHPLGVWANGSATHSGRRPNGYTLGL
GGGGSAQGSPRDVELPTQERQEGQEHPGLRGHLVQARVGASSWSSEFAPS
|
|
|
|---|
| BDBM50004566 |
|---|
| n/a |
|---|
| Name | BDBM50004566 |
|---|
| Synonyms: | 9-Chloro-2-furan-2-yl-[1,2,4]triazolo[1,5-c]quinazolin-5-ylamine | 9-Chloro-2-furan-2-yl-[1,2,4]triazolo[1,5-c]quinazolin-5-ylamine(CGS 15943) | 9-Chloro-[1,2,4]triazolo[1,5-c]quinazolin-5-ylamine(CGS 15943) | 9-chloro-2-(2-furanyl)-1,2,4-triazolo[1.5-c]quinazolin-5-amine | 9-chloro-2-(furan-2-yl)-[1,2,4]triazolo[1,5-c]quinazolin-5-amine | CGS-15943 | CHEMBL16687 | CHEMBL268431 | Nonnucleoside analog, 4 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C13H8ClN5O |
|---|
| Mol. Mass. | 285.689 |
|---|
| SMILES | Nc1nc2ccc(Cl)cc2c2nc(nn12)-c1ccco1 |
|---|
| Structure | 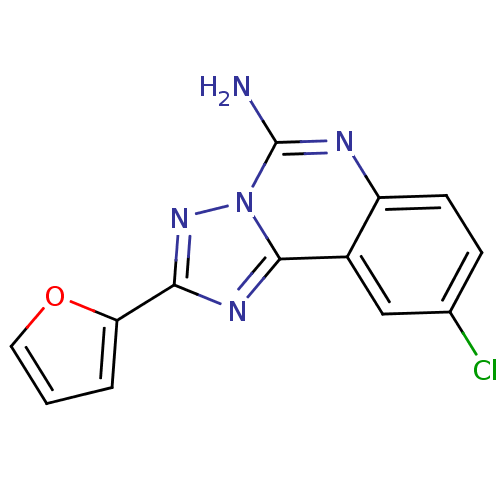
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Baraldi, PG; Cacciari, B; Spalluto, G; Pineda de las Infantas y Villatoro, MJ; Zocchi, C; Dionisotti, S; Ongini, E Pyrazolo[4,3-e]-1,2,4-triazolo[1,5-c]pyrimidine derivatives: potent and selective A(2A) adenosine antagonists. J Med Chem39:1164-71 (1996) [PubMed] Article
Baraldi, PG; Cacciari, B; Spalluto, G; Pineda de las Infantas y Villatoro, MJ; Zocchi, C; Dionisotti, S; Ongini, E Pyrazolo[4,3-e]-1,2,4-triazolo[1,5-c]pyrimidine derivatives: potent and selective A(2A) adenosine antagonists. J Med Chem39:1164-71 (1996) [PubMed] Article