| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Vitamin K-dependent protein C |
|---|
| Ligand | BDBM50114445 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEBML_27845 |
|---|
| IC50 | >11000±n/a nM |
|---|
| Citation |  Jia, ZJ; Wu, Y; Huang, W; Goldman, E; Zhang, P; Woolfrey, J; Wong, P; Huang, B; Sinha, U; Park, G; Reed, A; Scarborough, RM; Zhu, BY Design, synthesis and biological activity of novel non-amidine factor Xa inhibitors. Part 1: P(1) structure-activity relationships of the substituted 1-(2-Naphthyl)-1H-pyrazole-5-carboxylamides. Bioorg Med Chem Lett12:1651-5 (2002) [PubMed] Jia, ZJ; Wu, Y; Huang, W; Goldman, E; Zhang, P; Woolfrey, J; Wong, P; Huang, B; Sinha, U; Park, G; Reed, A; Scarborough, RM; Zhu, BY Design, synthesis and biological activity of novel non-amidine factor Xa inhibitors. Part 1: P(1) structure-activity relationships of the substituted 1-(2-Naphthyl)-1H-pyrazole-5-carboxylamides. Bioorg Med Chem Lett12:1651-5 (2002) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Vitamin K-dependent protein C |
|---|
| Name: | Vitamin K-dependent protein C |
|---|
| Synonyms: | Activated protein C cofactor | Anticoagulant protein C | Apolipoprotein H | Autoprothrombin IIA | Blood coagulation factor XIV | Coagulation factor V | Coagulation factor V heavy chain | Coagulation factor V light chain | Endothelial protein C receptor | PROC | PROC_HUMAN | Proaccelerin, labile factor | Vitamin K-dependent protein C precursor |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 52067.73 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | n/a |
|---|
| Residue: | 461 |
|---|
| Sequence: | MWQLTSLLLFVATWGISGTPAPLDSVFSSSERAHQVLRIRKRANSFLEELRHSSLERECI
EEICDFEEAKEIFQNVDDTLAFWSKHVDGDQCLVLPLEHPCASLCCGHGTCIDGIGSFSC
DCRSGWEGRFCQREVSFLNCSLDNGGCTHYCLEEVGWRRCSCAPGYKLGDDLLQCHPAVK
FPCGRPWKRMEKKRSHLKRDTEDQEDQVDPRLIDGKMTRRGDSPWQVVLLDSKKKLACGA
VLIHPSWVLTAAHCMDESKKLLVRLGEYDLRRWEKWELDLDIKEVFVHPNYSKSTTDNDI
ALLHLAQPATLSQTIVPICLPDSGLAERELNQAGQETLVTGWGYHSSREKEAKRNRTFVL
NFIKIPVVPHNECSEVMSNMVSENMLCAGILGDRQDACEGDSGGPMVASFHGTWFLVGLV
SWGEGCGLLHNYGVYTKVSRYLDWIHGHIRDKEAPQKSWAP
|
|
|
|---|
| BDBM50114445 |
|---|
| n/a |
|---|
| Name | BDBM50114445 |
|---|
| Synonyms: | 2-(6-Hydroxy-naphthalen-2-yl)-5-methyl-2H-pyrazole-3-carboxylic acid (2'-sulfamoyl-biphenyl-4-yl)-amide | CHEMBL44753 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C27H22N4O4S |
|---|
| Mol. Mass. | 498.553 |
|---|
| SMILES | Cc1cc(C(=O)Nc2ccc(cc2)-c2ccccc2S(N)(=O)=O)n(n1)-c1ccc2cc(O)ccc2c1 |
|---|
| Structure | 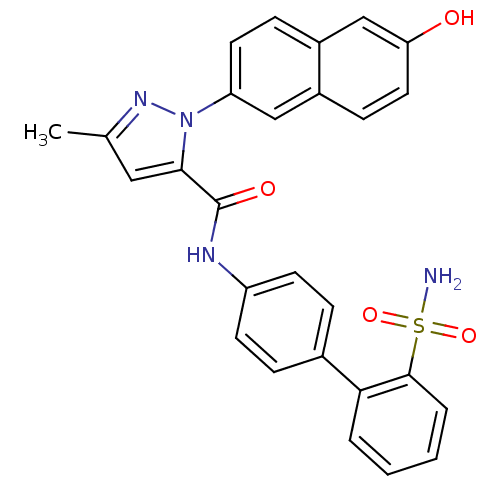
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Jia, ZJ; Wu, Y; Huang, W; Goldman, E; Zhang, P; Woolfrey, J; Wong, P; Huang, B; Sinha, U; Park, G; Reed, A; Scarborough, RM; Zhu, BY Design, synthesis and biological activity of novel non-amidine factor Xa inhibitors. Part 1: P(1) structure-activity relationships of the substituted 1-(2-Naphthyl)-1H-pyrazole-5-carboxylamides. Bioorg Med Chem Lett12:1651-5 (2002) [PubMed]
Jia, ZJ; Wu, Y; Huang, W; Goldman, E; Zhang, P; Woolfrey, J; Wong, P; Huang, B; Sinha, U; Park, G; Reed, A; Scarborough, RM; Zhu, BY Design, synthesis and biological activity of novel non-amidine factor Xa inhibitors. Part 1: P(1) structure-activity relationships of the substituted 1-(2-Naphthyl)-1H-pyrazole-5-carboxylamides. Bioorg Med Chem Lett12:1651-5 (2002) [PubMed]