| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Cytochrome P450 2D6 |
|---|
| Ligand | BDBM50161764 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_369007 (CHEMBL870289) |
|---|
| IC50 | >3000±n/a nM |
|---|
| Citation |  Wishka, DG; Walker, DP; Yates, KM; Reitz, SC; Jia, S; Myers, JK; Olson, KL; Jacobsen, EJ; Wolfe, ML; Groppi, VE; Hanchar, AJ; Thornburgh, BA; Cortes-Burgos, LA; Wong, EH; Staton, BA; Raub, TJ; Higdon, NR; Wall, TM; Hurst, RS; Walters, RR; Hoffmann, WE; Hajos, M; Franklin, S; Carey, G; Gold, LH; Cook, KK; Sands, SB; Zhao, SX; Soglia, JR; Kalgutkar, AS; Arneric, SP; Rogers, BN Discovery of N-[(3R)-1-azabicyclo[2.2.2]oct-3-yl]furo[2,3-c]pyridine-5-carboxamide, an agonist of the alpha7 nicotinic acetylcholine receptor, for the potential treatment of cognitive deficits in schizophrenia: synthesis and structure--activity relationship. J Med Chem49:4425-36 (2006) [PubMed] Article Wishka, DG; Walker, DP; Yates, KM; Reitz, SC; Jia, S; Myers, JK; Olson, KL; Jacobsen, EJ; Wolfe, ML; Groppi, VE; Hanchar, AJ; Thornburgh, BA; Cortes-Burgos, LA; Wong, EH; Staton, BA; Raub, TJ; Higdon, NR; Wall, TM; Hurst, RS; Walters, RR; Hoffmann, WE; Hajos, M; Franklin, S; Carey, G; Gold, LH; Cook, KK; Sands, SB; Zhao, SX; Soglia, JR; Kalgutkar, AS; Arneric, SP; Rogers, BN Discovery of N-[(3R)-1-azabicyclo[2.2.2]oct-3-yl]furo[2,3-c]pyridine-5-carboxamide, an agonist of the alpha7 nicotinic acetylcholine receptor, for the potential treatment of cognitive deficits in schizophrenia: synthesis and structure--activity relationship. J Med Chem49:4425-36 (2006) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Cytochrome P450 2D6 |
|---|
| Name: | Cytochrome P450 2D6 |
|---|
| Synonyms: | CP2D6_HUMAN | CYP2D6 | CYP2DL1 | CYPIID6 | Cytochrome P450 2D6 (CYP2D6) | Debrisoquine 4-hydroxylase | P450-DB1 |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 55774.82 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P10635 |
|---|
| Residue: | 497 |
|---|
| Sequence: | MGLEALVPLAVIVAIFLLLVDLMHRRQRWAARYPPGPLPLPGLGNLLHVDFQNTPYCFDQ
LRRRFGDVFSLQLAWTPVVVLNGLAAVREALVTHGEDTADRPPVPITQILGFGPRSQGVF
LARYGPAWREQRRFSVSTLRNLGLGKKSLEQWVTEEAACLCAAFANHSGRPFRPNGLLDK
AVSNVIASLTCGRRFEYDDPRFLRLLDLAQEGLKEESGFLREVLNAVPVLLHIPALAGKV
LRFQKAFLTQLDELLTEHRMTWDPAQPPRDLTEAFLAEMEKAKGNPESSFNDENLRIVVA
DLFSAGMVTTSTTLAWGLLLMILHPDVQRRVQQEIDDVIGQVRRPEMGDQAHMPYTTAVI
HEVQRFGDIVPLGVTHMTSRDIEVQGFRIPKGTTLITNLSSVLKDEAVWEKPFRFHPEHF
LDAQGHFVKPEAFLPFSAGRRACLGEPLARMELFLFFTSLLQHFSFSVPTGQPRPSHHGV
FAFLVSPSPYELCAVPR
|
|
|
|---|
| BDBM50161764 |
|---|
| n/a |
|---|
| Name | BDBM50161764 |
|---|
| Synonyms: | (R)-4-chloro-N-(quinuclidin-3-yl)benzamide | (R)-4-chloro-N-(quinuclidin-8-yl)benzamide | CHEMBL177611 | N-(R)-1-Aza-bicyclo[2.2.2]oct-3-yl-4-chloro-benzamide | PNU-282987 | PNU282987 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C14H17ClN2O |
|---|
| Mol. Mass. | 264.751 |
|---|
| SMILES | Clc1ccc(cc1)C(=O)N[C@H]1CN2CCC1CC2 |r,wU:10.10,TLB:9:10:14.13:16.17,(9.34,-31.61,;10.67,-32.38,;10.67,-33.92,;12,-34.69,;13.35,-33.92,;13.34,-32.37,;12,-31.6,;14.68,-34.69,;16.01,-33.92,;14.68,-36.23,;16.02,-37,;16.46,-35.89,;16.53,-37.53,;15.18,-38.13,;14.9,-39.53,;16.27,-38.89,;17.81,-39.55,;18,-38.17,)| |
|---|
| Structure | 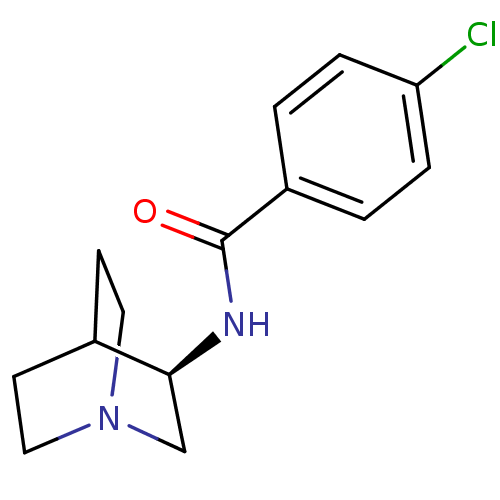
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Wishka, DG; Walker, DP; Yates, KM; Reitz, SC; Jia, S; Myers, JK; Olson, KL; Jacobsen, EJ; Wolfe, ML; Groppi, VE; Hanchar, AJ; Thornburgh, BA; Cortes-Burgos, LA; Wong, EH; Staton, BA; Raub, TJ; Higdon, NR; Wall, TM; Hurst, RS; Walters, RR; Hoffmann, WE; Hajos, M; Franklin, S; Carey, G; Gold, LH; Cook, KK; Sands, SB; Zhao, SX; Soglia, JR; Kalgutkar, AS; Arneric, SP; Rogers, BN Discovery of N-[(3R)-1-azabicyclo[2.2.2]oct-3-yl]furo[2,3-c]pyridine-5-carboxamide, an agonist of the alpha7 nicotinic acetylcholine receptor, for the potential treatment of cognitive deficits in schizophrenia: synthesis and structure--activity relationship. J Med Chem49:4425-36 (2006) [PubMed] Article
Wishka, DG; Walker, DP; Yates, KM; Reitz, SC; Jia, S; Myers, JK; Olson, KL; Jacobsen, EJ; Wolfe, ML; Groppi, VE; Hanchar, AJ; Thornburgh, BA; Cortes-Burgos, LA; Wong, EH; Staton, BA; Raub, TJ; Higdon, NR; Wall, TM; Hurst, RS; Walters, RR; Hoffmann, WE; Hajos, M; Franklin, S; Carey, G; Gold, LH; Cook, KK; Sands, SB; Zhao, SX; Soglia, JR; Kalgutkar, AS; Arneric, SP; Rogers, BN Discovery of N-[(3R)-1-azabicyclo[2.2.2]oct-3-yl]furo[2,3-c]pyridine-5-carboxamide, an agonist of the alpha7 nicotinic acetylcholine receptor, for the potential treatment of cognitive deficits in schizophrenia: synthesis and structure--activity relationship. J Med Chem49:4425-36 (2006) [PubMed] Article