| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Acetylcholinesterase |
|---|
| Ligand | BDBM50199515 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_441060 (CHEMBL890218) |
|---|
| pH | 5±n/a |
|---|
| IC50 | 78.4±n/a nM |
|---|
| Comments | extracted |
|---|
| Citation |  He, XC; Feng, S; Wang, ZF; Shi, Y; Zheng, S; Xia, Y; Jiang, H; Tang, XC; Bai, D Study on dual-site inhibitors of acetylcholinesterase: Highly potent derivatives of bis- and bifunctional huperzine B. Bioorg Med Chem15:1394-408 (2007) [PubMed] Article He, XC; Feng, S; Wang, ZF; Shi, Y; Zheng, S; Xia, Y; Jiang, H; Tang, XC; Bai, D Study on dual-site inhibitors of acetylcholinesterase: Highly potent derivatives of bis- and bifunctional huperzine B. Bioorg Med Chem15:1394-408 (2007) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Acetylcholinesterase |
|---|
| Name: | Acetylcholinesterase |
|---|
| Synonyms: | ACES_RAT | Acetylcholinesterase (AChE) | Acetylcholinesterase and butyrylcholinesterase (AChE and BChE) | Acetylcholinesterase precursor | Acetylcholinesterase, AChE | Ache |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 68193.62 |
|---|
| Organism: | Rattus norvegicus (rat) |
|---|
| Description: | P37136 |
|---|
| Residue: | 614 |
|---|
| Sequence: | MRPPWYPLHTPSLASPLLFLLLSLLGGGARAEGREDPQLLVRVRGGQLRGIRLKAPGGPV
SAFLGIPFAEPPVGSRRFMPPEPKRPWSGILDATTFQNVCYQYVDTLYPGFEGTEMWNPN
RELSEDCLYLNVWTPYPRPTSPTPVLIWIYGGGFYSGASSLDVYDGRFLAQVEGTVLVSM
NYRVGTFGFLALPGSREAPGNVGLLDQRLALQWVQENIAAFGGDPMSVTLFGESAGAASV
GMHILSLPSRSLFHRAVLQSGTPNGPWATVSAGEARRRATLLARLVGCPPGGAGGNDTEL
ISCLRTRPAQDLVDHEWHVLPQESIFRFSFVPVVDGDFLSDTPDALINTGDFQDLQVLVG
VVKDEGSYFLVYGVPGFSKDNESLISRAQFLAGVRIGVPQASDLAAEAVVLHYTDWLHPE
DPAHLRDAMSAVVGDHNVVCPVAQLAGRLAAQGARVYAYIFEHRASTLTWPLWMGVPHGY
EIEFIFGLPLDPSLNYTVEERIFAQRLMQYWTNFARTGDPNDPRDSKSPRWPPYTTAAQQ
YVSLNLKPLEVRRGLRAQTCAFWNRFLPKLLSATDTLDEAERQWKAEFHRWSSYMVHWKN
QFDHYSKQERCSDL
|
|
|
|---|
| BDBM50199515 |
|---|
| n/a |
|---|
| Name | BDBM50199515 |
|---|
| Synonyms: | CHEMBL244027 | N,N'-dimethyl-N,N'-bis[2-(1-oxo-8,15-didehydrolycodino)ethyl]-1,6-hexanediamine |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C44H64N6O2 |
|---|
| Mol. Mass. | 709.018 |
|---|
| SMILES | CN(CCCCCCN(C)CCN1CCC[C@@H]2[C@@H]3Cc4[nH]c(=O)ccc4[C@]12CC(C)=C3)CCN1CCC[C@@H]2[C@@H]3Cc4[nH]c(=O)ccc4[C@]12CC(C)=C3 |c:32,57,THB:29:28:16:18.19.25,45:46:37:48.51.49,41:40:37:48.51.49| |
|---|
| Structure | 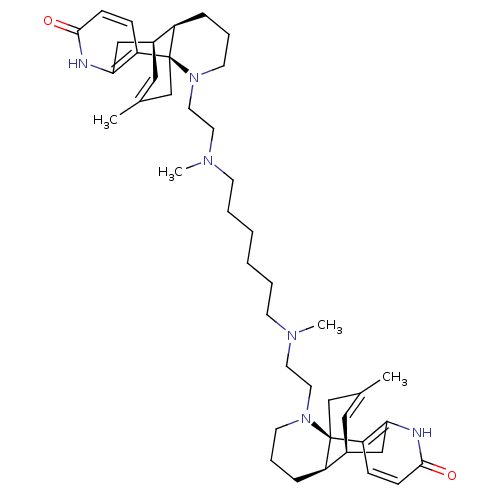
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 He, XC; Feng, S; Wang, ZF; Shi, Y; Zheng, S; Xia, Y; Jiang, H; Tang, XC; Bai, D Study on dual-site inhibitors of acetylcholinesterase: Highly potent derivatives of bis- and bifunctional huperzine B. Bioorg Med Chem15:1394-408 (2007) [PubMed] Article
He, XC; Feng, S; Wang, ZF; Shi, Y; Zheng, S; Xia, Y; Jiang, H; Tang, XC; Bai, D Study on dual-site inhibitors of acetylcholinesterase: Highly potent derivatives of bis- and bifunctional huperzine B. Bioorg Med Chem15:1394-408 (2007) [PubMed] Article