

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Glutamate receptor ionotropic, NMDA 2B | ||
| Ligand | BDBM50220609 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_457343 (CHEMBL940934) | ||
| Ki | >100±n/a nM | ||
| Citation |  Kawai, M; Sakurada, I; Morita, A; Iwamuro, Y; Ando, K; Omura, H; Sakakibara, S; Masuda, T; Koike, H; Honma, T; Hattori, K; Takashima, T; Mizuno, K; Mizutani, M; Kawamura, M Structure-activity relationship study of novel NR2B-selective antagonists with arylamides to avoid reactive metabolites formation. Bioorg Med Chem Lett17:5537-42 (2007) [PubMed] Article Kawai, M; Sakurada, I; Morita, A; Iwamuro, Y; Ando, K; Omura, H; Sakakibara, S; Masuda, T; Koike, H; Honma, T; Hattori, K; Takashima, T; Mizuno, K; Mizutani, M; Kawamura, M Structure-activity relationship study of novel NR2B-selective antagonists with arylamides to avoid reactive metabolites formation. Bioorg Med Chem Lett17:5537-42 (2007) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Glutamate receptor ionotropic, NMDA 2B | |||
| Name: | Glutamate receptor ionotropic, NMDA 2B | ||
| Synonyms: | GluN2B | Glutamate [NMDA] receptor subunit epsilon 2 | Grin2b | N-methyl D-aspartate receptor subtype 2B | NMDA receptor subunit N2B (GluN2B) | NMDAR2B | NMDE2_RAT | NR2B | ||
| Type: | Protein | ||
| Mol. Mass.: | 166077.66 | ||
| Organism: | Rattus norvegicus (Rat) | ||
| Description: | Q00960 | ||
| Residue: | 1482 | ||
| Sequence: |
| ||
| BDBM50220609 | |||
| n/a | |||
| Name | BDBM50220609 | ||
| Synonyms: | 3,5-dimethyl-N-(((1s,4s)-4-(2-phenoxyethyl)cyclohexyl)methyl)-1H-pyrazole-4-carboxamide | CHEMBL251496 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C21H29N3O2 | ||
| Mol. Mass. | 355.4739 | ||
| SMILES | Cc1n[nH]c(C)c1C(=O)NC[C@@H]1CC[C@H](CCOc2ccccc2)CC1 |wU:11.11,14.15,(19.66,-44.61,;19.98,-46.12,;18.95,-47.26,;19.73,-48.59,;21.23,-48.27,;22.38,-49.3,;21.39,-46.74,;22.72,-45.96,;22.72,-44.42,;24.06,-46.73,;25.39,-45.96,;26.72,-46.72,;26.72,-48.26,;28.05,-49.02,;29.38,-48.26,;30.72,-49.03,;32.05,-48.26,;33.38,-49.04,;34.72,-48.27,;36.05,-49.04,;37.38,-48.28,;37.39,-46.74,;36.05,-45.96,;34.72,-46.74,;29.38,-46.72,;28.05,-45.94,)| | ||
| Structure | 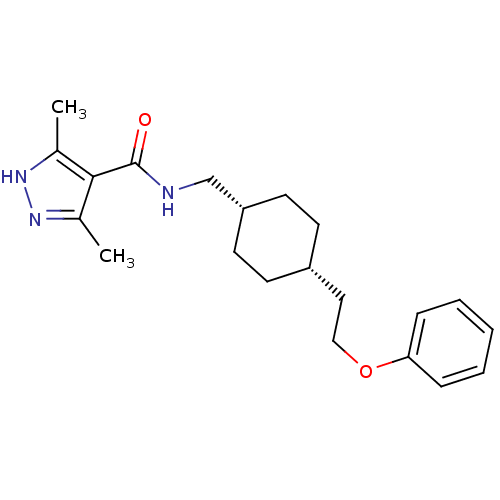
| ||