| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Matrix metalloproteinase-9 |
|---|
| Ligand | BDBM11551 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_461907 (CHEMBL944754) |
|---|
| Ki | 0.900000±n/a nM |
|---|
| Citation |  Yang, SM; Scannevin, RH; Wang, B; Burke, SL; Huang, Z; Karnachi, P; Wilson, LJ; Rhodes, KJ; Lagu, B; Murray, WV beta-N-Biaryl ether sulfonamide hydroxamates as potent gelatinase inhibitors: part 2. Optimization of alpha-amino substituents. Bioorg Med Chem Lett18:1140-5 (2008) [PubMed] Article Yang, SM; Scannevin, RH; Wang, B; Burke, SL; Huang, Z; Karnachi, P; Wilson, LJ; Rhodes, KJ; Lagu, B; Murray, WV beta-N-Biaryl ether sulfonamide hydroxamates as potent gelatinase inhibitors: part 2. Optimization of alpha-amino substituents. Bioorg Med Chem Lett18:1140-5 (2008) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Matrix metalloproteinase-9 |
|---|
| Name: | Matrix metalloproteinase-9 |
|---|
| Synonyms: | 67 kDa matrix metalloproteinase-9 | 82 kDa matrix metalloproteinase-9 | 92 kDa gelatinase | 92 kDa type IV collagenase | CLG4B | GELB | Gelatinase B | MMP-9 | MMP9 | MMP9_HUMAN | Matrix metalloproteinase 9 (MMP-9) | Matrix metalloproteinase-9 (MMP-9) | Matrix metalloproteinase-9 (MMP9) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 78452.28 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P14780 |
|---|
| Residue: | 707 |
|---|
| Sequence: | MSLWQPLVLVLLVLGCCFAAPRQRQSTLVLFPGDLRTNLTDRQLAEEYLYRYGYTRVAEM
RGESKSLGPALLLLQKQLSLPETGELDSATLKAMRTPRCGVPDLGRFQTFEGDLKWHHHN
ITYWIQNYSEDLPRAVIDDAFARAFALWSAVTPLTFTRVYSRDADIVIQFGVAEHGDGYP
FDGKDGLLAHAFPPGPGIQGDAHFDDDELWSLGKGVVVPTRFGNADGAACHFPFIFEGRS
YSACTTDGRSDGLPWCSTTANYDTDDRFGFCPSERLYTQDGNADGKPCQFPFIFQGQSYS
ACTTDGRSDGYRWCATTANYDRDKLFGFCPTRADSTVMGGNSAGELCVFPFTFLGKEYST
CTSEGRGDGRLWCATTSNFDSDKKWGFCPDQGYSLFLVAAHEFGHALGLDHSSVPEALMY
PMYRFTEGPPLHKDDVNGIRHLYGPRPEPEPRPPTTTTPQPTAPPTVCPTGPPTVHPSER
PTAGPTGPPSAGPTGPPTAGPSTATTVPLSPVDDACNVNIFDAIAEIGNQLYLFKDGKYW
RFSEGRGSRPQGPFLIADKWPALPRKLDSVFEERLSKKLFFFSGRQVWVYTGASVLGPRR
LDKLGLGADVAQVTGALRSGRGKMLLFSGRRLWRFDVKAQMVDPRSASEVDRMFPGVPLD
THDVFQYREKAYFCQDRFYWRVSSRSELNQVDQVGYVTYDILQCPED
|
|
|
|---|
| BDBM11551 |
|---|
| n/a |
|---|
| Name | BDBM11551 |
|---|
| Synonyms: | (3R)-N-hydroxy-2-[(4-methoxy-1,1-biphenyl-4-yl)methyl]-1,2-thiazinane-3-carboxamide 1,1-dioxide | (3R)-N-hydroxy-2-{[4-(4-methoxyphenyl)phenyl]methyl}-1,1-dioxo-1,2-thiazinane-3-carboxamide | CHEMBL103418 | Sultam Hydroxamate 23a |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C19H22N2O5S |
|---|
| Mol. Mass. | 390.453 |
|---|
| SMILES | COc1ccc(cc1)-c1ccc(CN2[C@H](CCCS2(=O)=O)C(=O)NO)cc1 |r| |
|---|
| Structure | 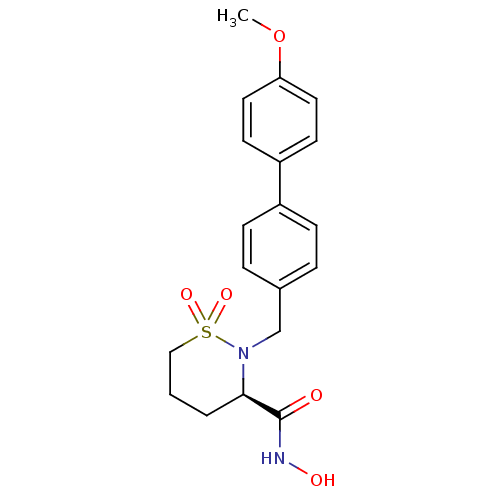
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Yang, SM; Scannevin, RH; Wang, B; Burke, SL; Huang, Z; Karnachi, P; Wilson, LJ; Rhodes, KJ; Lagu, B; Murray, WV beta-N-Biaryl ether sulfonamide hydroxamates as potent gelatinase inhibitors: part 2. Optimization of alpha-amino substituents. Bioorg Med Chem Lett18:1140-5 (2008) [PubMed] Article
Yang, SM; Scannevin, RH; Wang, B; Burke, SL; Huang, Z; Karnachi, P; Wilson, LJ; Rhodes, KJ; Lagu, B; Murray, WV beta-N-Biaryl ether sulfonamide hydroxamates as potent gelatinase inhibitors: part 2. Optimization of alpha-amino substituents. Bioorg Med Chem Lett18:1140-5 (2008) [PubMed] Article