| Reaction Details |
|---|
 | Report a problem with these data |
| Target | cAMP-dependent protein kinase catalytic subunit alpha |
|---|
| Ligand | BDBM3033 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEBML_161731 |
|---|
| IC50 | >100000±n/a nM |
|---|
| Citation |  Kleinschroth, J; Hartenstein, J; Rudolph, C; Schächtele, C Non-glycosidic/non-aminoalkyl-substituted indolocarbazoles as inhibitors of protein kinase C Bioorg Med Chem Lett3:1959-1964 (1993) Article Kleinschroth, J; Hartenstein, J; Rudolph, C; Schächtele, C Non-glycosidic/non-aminoalkyl-substituted indolocarbazoles as inhibitors of protein kinase C Bioorg Med Chem Lett3:1959-1964 (1993) Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| cAMP-dependent protein kinase catalytic subunit alpha |
|---|
| Name: | cAMP-dependent protein kinase catalytic subunit alpha |
|---|
| Synonyms: | KAPCA_BOVIN | PKA C-alpha | PKC | PRKACA | Protein Kinase C | Protein Kinase C, bovine brain | cAMP-Dependent Protein Kinase (PKA) | cAMP-dependent Protein Kinase, bovine heart | cAMP-dependent protein kinase A | cAMP-dependent protein kinase alpha-catalytic subunit | cAMP-dependent protein kinase, alpha-catalytic subunit |
|---|
| Type: | Enzyme Complex |
|---|
| Mol. Mass.: | 40627.77 |
|---|
| Organism: | Bos taurus (bovine) |
|---|
| Description: | The PKA holoenzyme purified from bovine heart, exists as an inactive tetrameric complex, which consists of a regulatory dimer associated with two catalytic subunits. It requires cAMP to activate the enzymatic reaction. |
|---|
| Residue: | 351 |
|---|
| Sequence: | MGNAAAAKKGSEQESVKEFLAKAKEDFLKKWENPAQNTAHLDQFERIKTLGTGSFGRVML
VKHMETGNHYAMKILDKQKVVKLKQIEHTLNEKRILQAVNFPFLVKLEFSFKDNSNLYMV
MEYVPGGEMFSHLRRIGRFSEPHARFYAAQIVLTFEYLHSLDLIYRDLKPENLLIDQQGY
IQVTDFGFAKRVKGRTWTLCGTPEYLAPEIILSKGYNKAVDWWALGVLIYEMAAGYPPFF
ADQPIQIYEKIVSGKVRFPSHFSSDLKDLLRNLLQVDLTKRFGNLKNGVNDIKNHKWFAT
TDWIAIYQRKVEAPFIPKFKGPGDTSNFDDYEEEEIRVSINEKCGKEFSEF
|
|
|
|---|
| BDBM3033 |
|---|
| n/a |
|---|
| Name | BDBM3033 |
|---|
| Synonyms: | 3-{23-methyl-14-oxo-3,13,23-triazahexacyclo[14.7.0.0^{2,10}.0^{4,9}.0^{11,15}.0^{17,22}]tricosa-1(16),2(10),4(9),5,7,11(15),17,19,21-nonaen-3-yl}propanenitrile | CHEMBL302449 | Go 6976 | Indolocarbazole deriv. |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C24H18N4O |
|---|
| Mol. Mass. | 378.4259 |
|---|
| SMILES | Cn1c2ccccc2c2c3C(=O)NCc3c3c4ccccc4n(CCC#N)c3c12 |
|---|
| Structure | 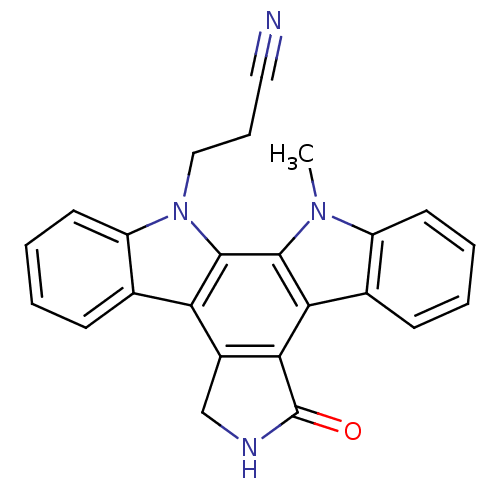
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Kleinschroth, J; Hartenstein, J; Rudolph, C; Schächtele, C Non-glycosidic/non-aminoalkyl-substituted indolocarbazoles as inhibitors of protein kinase C Bioorg Med Chem Lett3:1959-1964 (1993) Article
Kleinschroth, J; Hartenstein, J; Rudolph, C; Schächtele, C Non-glycosidic/non-aminoalkyl-substituted indolocarbazoles as inhibitors of protein kinase C Bioorg Med Chem Lett3:1959-1964 (1993) Article