| Reaction Details |
|---|
 | Report a problem with these data |
| Target | ATP-dependent molecular chaperone HSP82 |
|---|
| Ligand | BDBM50296578 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_582198 (CHEMBL1059858) |
|---|
| IC50 | 15400±n/a nM |
|---|
| Citation |  Hong, TJ; Park, H; Kim, YJ; Jeong, JH; Hahn, JS Identification of new Hsp90 inhibitors by structure-based virtual screening. Bioorg Med Chem Lett19:4839-42 (2009) [PubMed] Article Hong, TJ; Park, H; Kim, YJ; Jeong, JH; Hahn, JS Identification of new Hsp90 inhibitors by structure-based virtual screening. Bioorg Med Chem Lett19:4839-42 (2009) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| ATP-dependent molecular chaperone HSP82 |
|---|
| Name: | ATP-dependent molecular chaperone HSP82 |
|---|
| Synonyms: | 82 kDa heat shock protein | ATP-dependent molecular chaperone HSP82 | HSP82 | HSP82_YEAST | HSP90 | Heat Shock Protein 90 (Hsp90) | Heat shock protein Hsp90 heat-inducible isoform |
|---|
| Type: | Molecular Chaperone |
|---|
| Mol. Mass.: | 81369.08 |
|---|
| Organism: | Saccharomyces cerevisiae |
|---|
| Description: | Histidine-tagged yeast HSP90 was transformed into E. coli and purified (>90%) by metal affinity, gel filtration, and ion-exchange chromatography. |
|---|
| Residue: | 709 |
|---|
| Sequence: | MASETFEFQAEITQLMSLIINTVYSNKEIFLRELISNASDALDKIRYKSLSDPKQLETEP
DLFIRITPKPEQKVLEIRDSGIGMTKAELINNLGTIAKSGTKAFMEALSAGADVSMIGQF
GVGFYSLFLVADRVQVISKSNDDEQYIWESNAGGSFTVTLDEVNERIGRGTILRLFLKDD
QLEYLEEKRIKEVIKRHSEFVAYPIQLVVTKEVEKEVPIPEEEKKDEEKKDEEKKDEDDK
KPKLEEVDEEEEKKPKTKKVKEEVQEIEELNKTKPLWTRNPSDITQEEYNAFYKSISNDW
EDPLYVKHFSVEGQLEFRAILFIPKRAPFDLFESKKKKNNIKLYVRRVFITDEAEDLIPE
WLSFVKGVVDSEDLPLNLSREMLQQNKIMKVIRKNIVKKLIEAFNEIAEDSEQFEKFYSA
FSKNIKLGVHEDTQNRAALAKLLRYNSTKSVDELTSLTDYVTRMPEHQKNIYYITGESLK
AVEKSPFLDALKAKNFEVLFLTDPIDEYAFTQLKEFEGKTLVDITKDFELEETDEEKAER
EKEIKEYEPLTKALKEILGDQVEKVVVSYKLLDAPAAIRTGQFGWSANMERIMKAQALRD
SSMSSYMSSKKTFEISPKSPIIKELKKRVDEGGAQDKTVKDLTKLLYETALLTSGFSLDE
PTSFASRINRLISLGLNIDEDEETETAPEASTAAPVEEVPADTEMEEVD
|
|
|
|---|
| BDBM50296578 |
|---|
| n/a |
|---|
| Name | BDBM50296578 |
|---|
| Synonyms: | 2-(5-(benzo[d][1,3]dioxol-5-yl)-4-phenyl-4H-1,2,4-triazol-3-ylthio)-1-phenylethanone | CHEMBL564708 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C23H17N3O3S |
|---|
| Mol. Mass. | 415.464 |
|---|
| SMILES | O=C(CSc1nnc(-c2ccc3OCOc3c2)n1-c1ccccc1)c1ccccc1 |
|---|
| Structure | 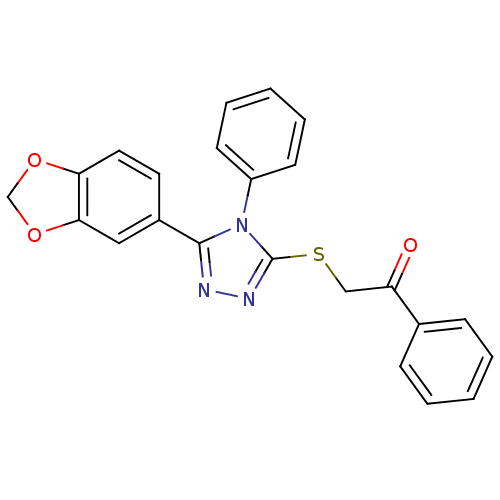
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Hong, TJ; Park, H; Kim, YJ; Jeong, JH; Hahn, JS Identification of new Hsp90 inhibitors by structure-based virtual screening. Bioorg Med Chem Lett19:4839-42 (2009) [PubMed] Article
Hong, TJ; Park, H; Kim, YJ; Jeong, JH; Hahn, JS Identification of new Hsp90 inhibitors by structure-based virtual screening. Bioorg Med Chem Lett19:4839-42 (2009) [PubMed] Article