| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Mitogen-activated protein kinase 9 |
|---|
| Ligand | BDBM50305006 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_606947 (CHEMBL1072690) |
|---|
| IC50 | 4100±n/a nM |
|---|
| Citation |  Harris, CM; Ericsson, AM; Argiriadi, MA; Barberis, C; Borhani, DW; Burchat, A; Calderwood, DJ; Cunha, GA; Dixon, RW; Frank, KE; Johnson, EF; Kamens, J; Kwak, S; Li, B; Mullen, KD; Perron, DC; Wang, L; Wishart, N; Wu, X; Zhang, X; Zmetra, TR; Talanian, RV 2,4-Diaminopyrimidine MK2 inhibitors. Part II: Structure-based inhibitor optimization. Bioorg Med Chem Lett20:334-7 (2010) [PubMed] Article Harris, CM; Ericsson, AM; Argiriadi, MA; Barberis, C; Borhani, DW; Burchat, A; Calderwood, DJ; Cunha, GA; Dixon, RW; Frank, KE; Johnson, EF; Kamens, J; Kwak, S; Li, B; Mullen, KD; Perron, DC; Wang, L; Wishart, N; Wu, X; Zhang, X; Zmetra, TR; Talanian, RV 2,4-Diaminopyrimidine MK2 inhibitors. Part II: Structure-based inhibitor optimization. Bioorg Med Chem Lett20:334-7 (2010) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Mitogen-activated protein kinase 9 |
|---|
| Name: | Mitogen-activated protein kinase 9 |
|---|
| Synonyms: | JNK-55 | JNK2 | JNK2/JNK3 | MAPK9 | MK09_HUMAN | Mitogen-Activated Protein Kinase 9 (JNK2) | Mitogen-activated protein kinase 8/9 | PRKM9 | SAPK1A | Stress-activated protein kinase JNK2 | c-Jun N-terminal kinase 2 | c-Jun N-terminal kinase 2 (JNK2) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 48131.49 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | JNK-2 was purchased from Upstate Cell Signaling Solutions (formerly Upstate Biotechnology). |
|---|
| Residue: | 424 |
|---|
| Sequence: | MSDSKCDSQFYSVQVADSTFTVLKRYQQLKPIGSGAQGIVCAAFDTVLGINVAVKKLSRP
FQNQTHAKRAYRELVLLKCVNHKNIISLLNVFTPQKTLEEFQDVYLVMELMDANLCQVIH
MELDHERMSYLLYQMLCGIKHLHSAGIIHRDLKPSNIVVKSDCTLKILDFGLARTACTNF
MMTPYVVTRYYRAPEVILGMGYKENVDIWSVGCIMGELVKGCVIFQGTDHIDQWNKVIEQ
LGTPSAEFMKKLQPTVRNYVENRPKYPGIKFEELFPDWIFPSESERDKIKTSQARDLLSK
MLVIDPDKRISVDEALRHPYITVWYDPAEAEAPPPQIYDAQLEEREHAIEEWKELIYKEV
MDWEERSKNGVVKDQPSDAAVSSNATPSQSSSINDISSMSTEQTLASDTDSSLDASTGPL
EGCR
|
|
|
|---|
| BDBM50305006 |
|---|
| n/a |
|---|
| Name | BDBM50305006 |
|---|
| Synonyms: | 5-(2-aminoethyl)-7-(7-(benzo[b]thiophen-2-yl)-1H-indazol-5-yl)-5H-pyrrolo[3,2-d]pyrimidin-2-amine | CHEMBL590109 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C23H19N7S |
|---|
| Mol. Mass. | 425.509 |
|---|
| SMILES | NCCn1cc(-c2cc(-c3cc4ccccc4s3)c3[nH]ncc3c2)c2nc(N)ncc12 |
|---|
| Structure | 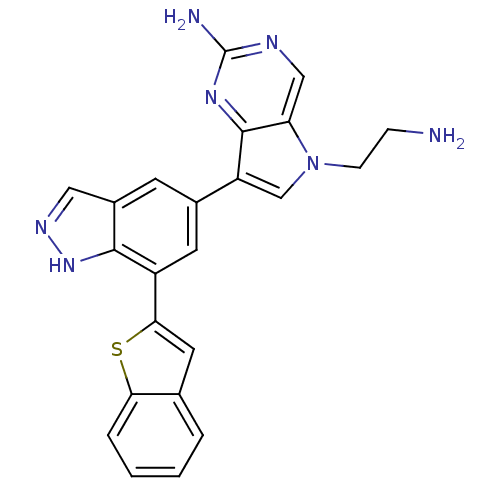
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Harris, CM; Ericsson, AM; Argiriadi, MA; Barberis, C; Borhani, DW; Burchat, A; Calderwood, DJ; Cunha, GA; Dixon, RW; Frank, KE; Johnson, EF; Kamens, J; Kwak, S; Li, B; Mullen, KD; Perron, DC; Wang, L; Wishart, N; Wu, X; Zhang, X; Zmetra, TR; Talanian, RV 2,4-Diaminopyrimidine MK2 inhibitors. Part II: Structure-based inhibitor optimization. Bioorg Med Chem Lett20:334-7 (2010) [PubMed] Article
Harris, CM; Ericsson, AM; Argiriadi, MA; Barberis, C; Borhani, DW; Burchat, A; Calderwood, DJ; Cunha, GA; Dixon, RW; Frank, KE; Johnson, EF; Kamens, J; Kwak, S; Li, B; Mullen, KD; Perron, DC; Wang, L; Wishart, N; Wu, X; Zhang, X; Zmetra, TR; Talanian, RV 2,4-Diaminopyrimidine MK2 inhibitors. Part II: Structure-based inhibitor optimization. Bioorg Med Chem Lett20:334-7 (2010) [PubMed] Article