| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Histone deacetylase 1 |
|---|
| Ligand | BDBM19428 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_625334 (CHEMBL1111708) |
|---|
| IC50 | 9±n/a nM |
|---|
| Citation |  Cho, YS; Whitehead, L; Li, J; Chen, CH; Jiang, L; Vögtle, M; Francotte, E; Richert, P; Wagner, T; Traebert, M; Lu, Q; Cao, X; Dumotier, B; Fejzo, J; Rajan, S; Wang, P; Yan-Neale, Y; Shao, W; Atadja, P; Shultz, M Conformational refinement of hydroxamate-based histone deacetylase inhibitors and exploration of 3-piperidin-3-ylindole analogues of dacinostat (LAQ824). J Med Chem53:2952-63 (2010) [PubMed] Article Cho, YS; Whitehead, L; Li, J; Chen, CH; Jiang, L; Vögtle, M; Francotte, E; Richert, P; Wagner, T; Traebert, M; Lu, Q; Cao, X; Dumotier, B; Fejzo, J; Rajan, S; Wang, P; Yan-Neale, Y; Shao, W; Atadja, P; Shultz, M Conformational refinement of hydroxamate-based histone deacetylase inhibitors and exploration of 3-piperidin-3-ylindole analogues of dacinostat (LAQ824). J Med Chem53:2952-63 (2010) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Histone deacetylase 1 |
|---|
| Name: | Histone deacetylase 1 |
|---|
| Synonyms: | Cereblon/Histone deacetylase 1 | HD1 | HDAC1 | HDAC1_HUMAN | Histone deacetylase 1 (HDAC1) | Human HDAC1 | RPD3L1 |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 55090.27 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q13547 |
|---|
| Residue: | 482 |
|---|
| Sequence: | MAQTQGTRRKVCYYYDGDVGNYYYGQGHPMKPHRIRMTHNLLLNYGLYRKMEIYRPHKAN
AEEMTKYHSDDYIKFLRSIRPDNMSEYSKQMQRFNVGEDCPVFDGLFEFCQLSTGGSVAS
AVKLNKQQTDIAVNWAGGLHHAKKSEASGFCYVNDIVLAILELLKYHQRVLYIDIDIHHG
DGVEEAFYTTDRVMTVSFHKYGEYFPGTGDLRDIGAGKGKYYAVNYPLRDGIDDESYEAI
FKPVMSKVMEMFQPSAVVLQCGSDSLSGDRLGCFNLTIKGHAKCVEFVKSFNLPMLMLGG
GGYTIRNVARCWTYETAVALDTEIPNELPYNDYFEYFGPDFKLHISPSNMTNQNTNEYLE
KIKQRLFENLRMLPHAPGVQMQAIPEDAIPEESGDEDEDDPDKRISICSSDKRIACEEEF
SDSEEEGEGGRKNSSNFKKAKRVKTEDEKEKDPEEKKEVTEEEKTKEEKPEAKGVKEEVK
LA
|
|
|
|---|
| BDBM19428 |
|---|
| n/a |
|---|
| Name | BDBM19428 |
|---|
| Synonyms: | (2E)-N-hydroxy-3-(4-{[(2-hydroxyethyl)[2-(1H-indol-3-yl)ethyl]amino]methyl}phenyl)prop-2-enamide | (E)-N-hydroxy-3-[4-[[2-hydroxyethyl-[2-(1H-indol-3-yl)ethyl]amino]methyl]phenyl]prop-2-enamide | CHEMBL356066 | Dacinostat | LAQ-824 | NVP-LAQ824 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C22H25N3O3 |
|---|
| Mol. Mass. | 379.4522 |
|---|
| SMILES | OCCN(CCc1c[nH]c2ccccc12)Cc1ccc(\C=C\C(=O)NO)cc1 |
|---|
| Structure | 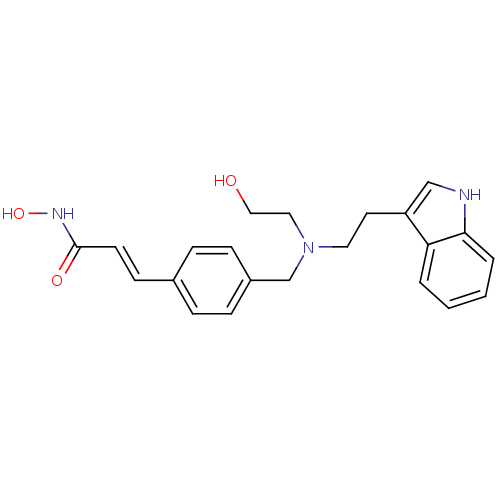
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Cho, YS; Whitehead, L; Li, J; Chen, CH; Jiang, L; Vögtle, M; Francotte, E; Richert, P; Wagner, T; Traebert, M; Lu, Q; Cao, X; Dumotier, B; Fejzo, J; Rajan, S; Wang, P; Yan-Neale, Y; Shao, W; Atadja, P; Shultz, M Conformational refinement of hydroxamate-based histone deacetylase inhibitors and exploration of 3-piperidin-3-ylindole analogues of dacinostat (LAQ824). J Med Chem53:2952-63 (2010) [PubMed] Article
Cho, YS; Whitehead, L; Li, J; Chen, CH; Jiang, L; Vögtle, M; Francotte, E; Richert, P; Wagner, T; Traebert, M; Lu, Q; Cao, X; Dumotier, B; Fejzo, J; Rajan, S; Wang, P; Yan-Neale, Y; Shao, W; Atadja, P; Shultz, M Conformational refinement of hydroxamate-based histone deacetylase inhibitors and exploration of 3-piperidin-3-ylindole analogues of dacinostat (LAQ824). J Med Chem53:2952-63 (2010) [PubMed] Article