| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Cathepsin D |
|---|
| Ligand | BDBM50328048 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_665268 (CHEMBL1260849) |
|---|
| IC50 | 8±n/a nM |
|---|
| Citation |  Probst, GD; Bowers, S; Sealy, JM; Stupi, B; Dressen, D; Jagodzinska, BM; Aquino, J; Gailunas, A; Truong, AP; Tso, L; Xu, YZ; Hom, RK; John, V; Tung, JS; Pleiss, MA; Tucker, JA; Konradi, AW; Sham, HL; Jagodzinski, J; Toth, G; Brecht, E; Yao, N; Pan, H; Lin, M; Artis, DR; Ruslim, L; Bova, MP; Sinha, S; Yednock, TA; Gauby, S; Zmolek, W; Quinn, KP; Sauer, JM Design and synthesis of hydroxyethylamine (HEA) BACE-1 inhibitors: structure-activity relationship of the aryl region. Bioorg Med Chem Lett20:6034-9 (2010) [PubMed] Article Probst, GD; Bowers, S; Sealy, JM; Stupi, B; Dressen, D; Jagodzinska, BM; Aquino, J; Gailunas, A; Truong, AP; Tso, L; Xu, YZ; Hom, RK; John, V; Tung, JS; Pleiss, MA; Tucker, JA; Konradi, AW; Sham, HL; Jagodzinski, J; Toth, G; Brecht, E; Yao, N; Pan, H; Lin, M; Artis, DR; Ruslim, L; Bova, MP; Sinha, S; Yednock, TA; Gauby, S; Zmolek, W; Quinn, KP; Sauer, JM Design and synthesis of hydroxyethylamine (HEA) BACE-1 inhibitors: structure-activity relationship of the aryl region. Bioorg Med Chem Lett20:6034-9 (2010) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Cathepsin D |
|---|
| Name: | Cathepsin D |
|---|
| Synonyms: | CATD_HUMAN | CPSD | CTSD | Cathepsin D [Precursor] | Cathepsin D heavy chain | Cathepsin D light chain | Cathepsin D precursor |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 44551.72 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Human proCathepsin D (SwissProt accession number P07339) was expressed in Sf9 cells, purified, and autoactivated. |
|---|
| Residue: | 412 |
|---|
| Sequence: | MQPSSLLPLALCLLAAPASALVRIPLHKFTSIRRTMSEVGGSVEDLIAKGPVSKYSQAVP
AVTEGPIPEVLKNYMDAQYYGEIGIGTPPQCFTVVFDTGSSNLWVPSIHCKLLDIACWIH
HKYNSDKSSTYVKNGTSFDIHYGSGSLSGYLSQDTVSVPCQSASSASALGGVKVERQVFG
EATKQPGITFIAAKFDGILGMAYPRISVNNVLPVFDNLMQQKLVDQNIFSFYLSRDPDAQ
PGGELMLGGTDSKYYKGSLSYLNVTRKAYWQVHLDQVEVASGLTLCKEGCEAIVDTGTSL
MVGPVDEVRELQKAIGAVPLIQGEYMIPCEKVSTLPAITLKLGGKGYKLSPEDYTLKVSQ
AGKTLCLSGFMGMDIPPPSGPLWILGDVFIGRYYTVFDRDNNRVGFAEAARL
|
|
|
|---|
| BDBM50328048 |
|---|
| n/a |
|---|
| Name | BDBM50328048 |
|---|
| Synonyms: | CHEMBL1257185 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C30H41F2N5O2 |
|---|
| Mol. Mass. | 541.6756 |
|---|
| SMILES | CC(=O)N[C@@H](Cc1cc(F)cc(F)c1)[C@H](O)CNC1(CCCCC1)c1cn(nn1)C12CC3CC(CC(C3)C1)C2 |r,TLB:26:29:32.31.36:34,THB:30:31:34:38.29.37,30:29:32.31.36:34,26:29:32:36.35.34,37:29:32:36.35.34,37:35:32:38.30.29| |
|---|
| Structure | 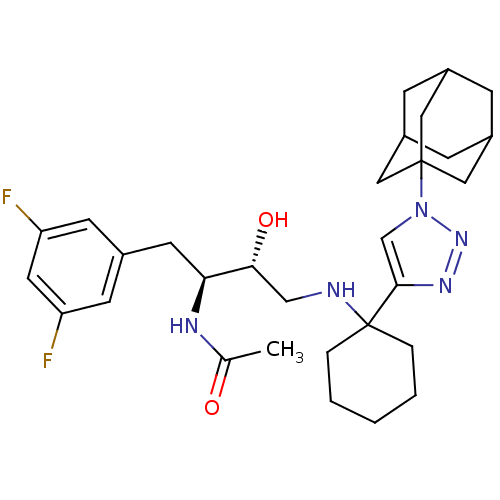
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Probst, GD; Bowers, S; Sealy, JM; Stupi, B; Dressen, D; Jagodzinska, BM; Aquino, J; Gailunas, A; Truong, AP; Tso, L; Xu, YZ; Hom, RK; John, V; Tung, JS; Pleiss, MA; Tucker, JA; Konradi, AW; Sham, HL; Jagodzinski, J; Toth, G; Brecht, E; Yao, N; Pan, H; Lin, M; Artis, DR; Ruslim, L; Bova, MP; Sinha, S; Yednock, TA; Gauby, S; Zmolek, W; Quinn, KP; Sauer, JM Design and synthesis of hydroxyethylamine (HEA) BACE-1 inhibitors: structure-activity relationship of the aryl region. Bioorg Med Chem Lett20:6034-9 (2010) [PubMed] Article
Probst, GD; Bowers, S; Sealy, JM; Stupi, B; Dressen, D; Jagodzinska, BM; Aquino, J; Gailunas, A; Truong, AP; Tso, L; Xu, YZ; Hom, RK; John, V; Tung, JS; Pleiss, MA; Tucker, JA; Konradi, AW; Sham, HL; Jagodzinski, J; Toth, G; Brecht, E; Yao, N; Pan, H; Lin, M; Artis, DR; Ruslim, L; Bova, MP; Sinha, S; Yednock, TA; Gauby, S; Zmolek, W; Quinn, KP; Sauer, JM Design and synthesis of hydroxyethylamine (HEA) BACE-1 inhibitors: structure-activity relationship of the aryl region. Bioorg Med Chem Lett20:6034-9 (2010) [PubMed] Article