| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Mitogen-activated protein kinase 8 |
|---|
| Ligand | BDBM50352631 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_767684 (CHEMBL1825796) |
|---|
| IC50 | 0.7±n/a nM |
|---|
| Citation |  Stebbins, JL; De, SK; Pavlickova, P; Chen, V; Machleidt, T; Chen, LH; Kuntzen, C; Kitada, S; Karin, M; Pellecchia, M Design and characterization of a potent and selective dual ATP- and substrate-competitive subnanomolar bidentate c-Jun N-terminal kinase (JNK) inhibitor. J Med Chem54:6206-14 (2011) [PubMed] Article Stebbins, JL; De, SK; Pavlickova, P; Chen, V; Machleidt, T; Chen, LH; Kuntzen, C; Kitada, S; Karin, M; Pellecchia, M Design and characterization of a potent and selective dual ATP- and substrate-competitive subnanomolar bidentate c-Jun N-terminal kinase (JNK) inhibitor. J Med Chem54:6206-14 (2011) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Mitogen-activated protein kinase 8 |
|---|
| Name: | Mitogen-activated protein kinase 8 |
|---|
| Synonyms: | JNK-46 | JNK1 | JNK1-alpha-1 | MAPK8 | MK08_HUMAN | Mitogen-Activated Protein Kinase 8 (JNK1) | PRKM8 | SAPK1 | SAPK1C | Stress-activated protein kinase JNK1 | c-Jun N-terminal kinase 1 | c-Jun N-terminal kinase 1 (JNK1) | c-Jun N-terminal kinase 1(JNK1) | c-Jun N-terminal kinase 2 (JNK2) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 48297.57 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | JNK-1 was purchased from Upstate Cell Signaling Solutions (formerly Upstate Biotechnology). |
|---|
| Residue: | 427 |
|---|
| Sequence: | MSRSKRDNNFYSVEIGDSTFTVLKRYQNLKPIGSGAQGIVCAAYDAILERNVAIKKLSRP
FQNQTHAKRAYRELVLMKCVNHKNIIGLLNVFTPQKSLEEFQDVYIVMELMDANLCQVIQ
MELDHERMSYLLYQMLCGIKHLHSAGIIHRDLKPSNIVVKSDCTLKILDFGLARTAGTSF
MMTPYVVTRYYRAPEVILGMGYKENVDLWSVGCIMGEMVCHKILFPGRDYIDQWNKVIEQ
LGTPCPEFMKKLQPTVRTYVENRPKYAGYSFEKLFPDVLFPADSEHNKLKASQARDLLSK
MLVIDASKRISVDEALQHPYINVWYDPSEAEAPPPKIPDKQLDEREHTIEEWKELIYKEV
MDLEERTKNGVIRGQPSPLGAAVINGSQHPSSSSSVNDVSSMSTDPTLASDTDSSLEAAA
GPLGCCR
|
|
|
|---|
| BDBM50352631 |
|---|
| n/a |
|---|
| Name | BDBM50352631 |
|---|
| Synonyms: | CHEMBL1822313 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C50H74N16O12 |
|---|
| Mol. Mass. | 1091.2226 |
|---|
| SMILES | CC(C)C[C@H](NC(=O)[C@@H](NC(=O)[C@@H](NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)CCCNC(N)=N)[C@@H](C)O)[C@@H](C)O)C(=O)N[C@@H](CC(N)=O)C(=O)NCC(=O)NCC(=O)NCCCNC(=O)c1ccc(cc1)-c1n[nH]c2ccccc12 |r| |
|---|
| Structure | 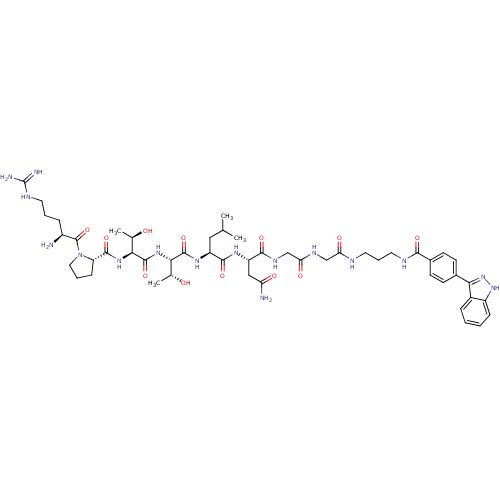
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Stebbins, JL; De, SK; Pavlickova, P; Chen, V; Machleidt, T; Chen, LH; Kuntzen, C; Kitada, S; Karin, M; Pellecchia, M Design and characterization of a potent and selective dual ATP- and substrate-competitive subnanomolar bidentate c-Jun N-terminal kinase (JNK) inhibitor. J Med Chem54:6206-14 (2011) [PubMed] Article
Stebbins, JL; De, SK; Pavlickova, P; Chen, V; Machleidt, T; Chen, LH; Kuntzen, C; Kitada, S; Karin, M; Pellecchia, M Design and characterization of a potent and selective dual ATP- and substrate-competitive subnanomolar bidentate c-Jun N-terminal kinase (JNK) inhibitor. J Med Chem54:6206-14 (2011) [PubMed] Article