| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Mitogen-activated protein kinase 9 |
|---|
| Ligand | BDBM50355393 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_773612 (CHEMBL1839978) |
|---|
| IC50 | >10000±n/a nM |
|---|
| Citation |  Guagnano, V; Furet, P; Spanka, C; Bordas, V; Le Douget, M; Stamm, C; Brueggen, J; Jensen, MR; Schnell, C; Schmid, H; Wartmann, M; Berghausen, J; Drueckes, P; Zimmerlin, A; Bussiere, D; Murray, J; Graus Porta, D Discovery of 3-(2,6-dichloro-3,5-dimethoxy-phenyl)-1-{6-[4-(4-ethyl-piperazin-1-yl)-phenylamino]-pyrimidin-4-yl}-1-methyl-urea (NVP-BGJ398), a potent and selective inhibitor of the fibroblast growth factor receptor family of receptor tyrosine kinase. J Med Chem54:7066-83 (2011) [PubMed] Article Guagnano, V; Furet, P; Spanka, C; Bordas, V; Le Douget, M; Stamm, C; Brueggen, J; Jensen, MR; Schnell, C; Schmid, H; Wartmann, M; Berghausen, J; Drueckes, P; Zimmerlin, A; Bussiere, D; Murray, J; Graus Porta, D Discovery of 3-(2,6-dichloro-3,5-dimethoxy-phenyl)-1-{6-[4-(4-ethyl-piperazin-1-yl)-phenylamino]-pyrimidin-4-yl}-1-methyl-urea (NVP-BGJ398), a potent and selective inhibitor of the fibroblast growth factor receptor family of receptor tyrosine kinase. J Med Chem54:7066-83 (2011) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Mitogen-activated protein kinase 9 |
|---|
| Name: | Mitogen-activated protein kinase 9 |
|---|
| Synonyms: | JNK-55 | JNK2 | JNK2/JNK3 | MAPK9 | MK09_HUMAN | Mitogen-Activated Protein Kinase 9 (JNK2) | Mitogen-activated protein kinase 8/9 | PRKM9 | SAPK1A | Stress-activated protein kinase JNK2 | c-Jun N-terminal kinase 2 | c-Jun N-terminal kinase 2 (JNK2) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 48131.49 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | JNK-2 was purchased from Upstate Cell Signaling Solutions (formerly Upstate Biotechnology). |
|---|
| Residue: | 424 |
|---|
| Sequence: | MSDSKCDSQFYSVQVADSTFTVLKRYQQLKPIGSGAQGIVCAAFDTVLGINVAVKKLSRP
FQNQTHAKRAYRELVLLKCVNHKNIISLLNVFTPQKTLEEFQDVYLVMELMDANLCQVIH
MELDHERMSYLLYQMLCGIKHLHSAGIIHRDLKPSNIVVKSDCTLKILDFGLARTACTNF
MMTPYVVTRYYRAPEVILGMGYKENVDIWSVGCIMGELVKGCVIFQGTDHIDQWNKVIEQ
LGTPSAEFMKKLQPTVRNYVENRPKYPGIKFEELFPDWIFPSESERDKIKTSQARDLLSK
MLVIDPDKRISVDEALRHPYITVWYDPAEAEAPPPQIYDAQLEEREHAIEEWKELIYKEV
MDWEERSKNGVVKDQPSDAAVSSNATPSQSSSINDISSMSTEQTLASDTDSSLDASTGPL
EGCR
|
|
|
|---|
| BDBM50355393 |
|---|
| n/a |
|---|
| Name | BDBM50355393 |
|---|
| Synonyms: | BGJ398 | CHEMBL1834657 | US9434697, BGJ398 | US9730931, BGJ398 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C26H31Cl2N7O3 |
|---|
| Mol. Mass. | 560.475 |
|---|
| SMILES | CCN1CCN(CC1)c1ccc(Nc2cc(ncn2)N(C)C(=O)Nc2c(Cl)c(OC)cc(OC)c2Cl)cc1 |
|---|
| Structure | 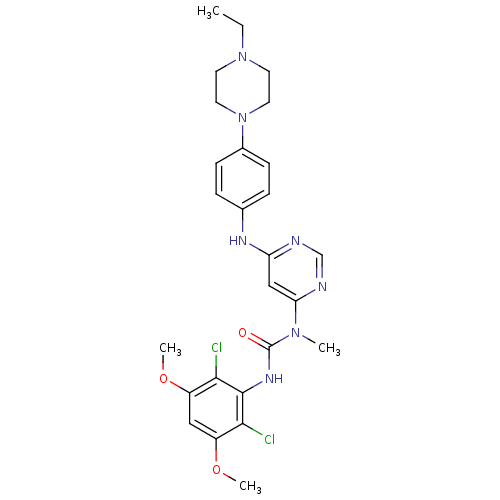
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Guagnano, V; Furet, P; Spanka, C; Bordas, V; Le Douget, M; Stamm, C; Brueggen, J; Jensen, MR; Schnell, C; Schmid, H; Wartmann, M; Berghausen, J; Drueckes, P; Zimmerlin, A; Bussiere, D; Murray, J; Graus Porta, D Discovery of 3-(2,6-dichloro-3,5-dimethoxy-phenyl)-1-{6-[4-(4-ethyl-piperazin-1-yl)-phenylamino]-pyrimidin-4-yl}-1-methyl-urea (NVP-BGJ398), a potent and selective inhibitor of the fibroblast growth factor receptor family of receptor tyrosine kinase. J Med Chem54:7066-83 (2011) [PubMed] Article
Guagnano, V; Furet, P; Spanka, C; Bordas, V; Le Douget, M; Stamm, C; Brueggen, J; Jensen, MR; Schnell, C; Schmid, H; Wartmann, M; Berghausen, J; Drueckes, P; Zimmerlin, A; Bussiere, D; Murray, J; Graus Porta, D Discovery of 3-(2,6-dichloro-3,5-dimethoxy-phenyl)-1-{6-[4-(4-ethyl-piperazin-1-yl)-phenylamino]-pyrimidin-4-yl}-1-methyl-urea (NVP-BGJ398), a potent and selective inhibitor of the fibroblast growth factor receptor family of receptor tyrosine kinase. J Med Chem54:7066-83 (2011) [PubMed] Article