| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform |
|---|
| Ligand | BDBM50357650 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_786051 (CHEMBL1919682) |
|---|
| IC50 | 9.6±n/a nM |
|---|
| Citation |  Nishimura, N; Siegmund, A; Liu, L; Yang, K; Bryan, MC; Andrews, KL; Bo, Y; Booker, SK; Caenepeel, S; Freeman, D; Liao, H; McCarter, J; Mullady, EL; San Miguel, T; Subramanian, R; Tamayo, N; Wang, L; Whittington, DA; Zalameda, L; Zhang, N; Hughes, PE; Norman, MH Phospshoinositide 3-kinase (PI3K)/mammalian target of rapamycin (mTOR) dual inhibitors: discovery and structure-activity relationships of a series of quinoline and quinoxaline derivatives. J Med Chem54:4735-51 (2011) [PubMed] Article Nishimura, N; Siegmund, A; Liu, L; Yang, K; Bryan, MC; Andrews, KL; Bo, Y; Booker, SK; Caenepeel, S; Freeman, D; Liao, H; McCarter, J; Mullady, EL; San Miguel, T; Subramanian, R; Tamayo, N; Wang, L; Whittington, DA; Zalameda, L; Zhang, N; Hughes, PE; Norman, MH Phospshoinositide 3-kinase (PI3K)/mammalian target of rapamycin (mTOR) dual inhibitors: discovery and structure-activity relationships of a series of quinoline and quinoxaline derivatives. J Med Chem54:4735-51 (2011) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform |
|---|
| Name: | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform |
|---|
| Synonyms: | PI3-kinase p110 subunit beta | PI3-kinase subunit p110-beta | PI3Kbeta | PIK3C1 | PIK3CB | PK3CB_HUMAN | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta (PI3Kbeta) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform (PI3K beta) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform (PI3K) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform (PI3K-beta) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform (PI3Kbeta) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform (PI3Kÿ²) | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit beta isoform | Phosphoinositide 3-Kinase (PI3K), beta | Phosphoinositide 3-Kinase (PI3K), beta Chain A | Phosphoinositide-3-kinase (PI3K beta) | PtdIns-3-kinase p110 |
|---|
| Type: | Enzyme Subunit |
|---|
| Mol. Mass.: | 122769.00 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P42338 |
|---|
| Residue: | 1070 |
|---|
| Sequence: | MCFSFIMPPAMADILDIWAVDSQIASDGSIPVDFLLPTGIYIQLEVPREATISYIKQMLW
KQVHNYPMFNLLMDIDSYMFACVNQTAVYEELEDETRRLCDVRPFLPVLKLVTRSCDPGE
KLDSKIGVLIGKGLHEFDSLKDPEVNEFRRKMRKFSEEKILSLVGLSWMDWLKQTYPPEH
EPSIPENLEDKLYGGKLIVAVHFENCQDVFSFQVSPNMNPIKVNELAIQKRLTIHGKEDE
VSPYDYVLQVSGRVEYVFGDHPLIQFQYIRNCVMNRALPHFILVECCKIKKMYEQEMIAI
EAAINRNSSNLPLPLPPKKTRIISHVWENNNPFQIVLVKGNKLNTEETVKVHVRAGLFHG
TELLCKTIVSSEVSGKNDHIWNEPLEFDINICDLPRMARLCFAVYAVLDKVKTKKSTKTI
NPSKYQTIRKAGKVHYPVAWVNTMVFDFKGQLRTGDIILHSWSSFPDELEEMLNPMGTVQ
TNPYTENATALHVKFPENKKQPYYYPPFDKIIEKAAEIASSDSANVSSRGGKKFLPVLKE
ILDRDPLSQLCENEMDLIWTLRQDCREIFPQSLPKLLLSIKWNKLEDVAQLQALLQIWPK
LPPREALELLDFNYPDQYVREYAVGCLRQMSDEELSQYLLQLVQVLKYEPFLDCALSRFL
LERALGNRRIGQFLFWHLRSEVHIPAVSVQFGVILEAYCRGSVGHMKVLSKQVEALNKLK
TLNSLIKLNAVKLNRAKGKEAMHTCLKQSAYREALSDLQSPLNPCVILSELYVEKCKYMD
SKMKPLWLVYNNKVFGEDSVGVIFKNGDDLRQDMLTLQMLRLMDLLWKEAGLDLRMLPYG
CLATGDRSGLIEVVSTSETIADIQLNSSNVAAAAAFNKDALLNWLKEYNSGDDLDRAIEE
FTLSCAGYCVASYVLGIGDRHSDNIMVKKTGQLFHIDFGHILGNFKSKFGIKRERVPFIL
TYDFIHVIQQGKTGNTEKFGRFRQCCEDAYLILRRHGNLFITLFALMLTAGLPELTSVKD
IQYLKDSLALGKSEEEALKQFKQKFDEALRESWTTKVNWMAHTVRKDYRS
|
|
|
|---|
| BDBM50357650 |
|---|
| n/a |
|---|
| Name | BDBM50357650 |
|---|
| Synonyms: | CHEMBL1914742 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C26H26ClFN6O2S |
|---|
| Mol. Mass. | 541.04 |
|---|
| SMILES | CC(C)N1CCN(CC1)c1cnc2ccc(cc2n1)-c1cnc(Cl)c(NS(=O)(=O)c2ccc(F)cc2)c1 |
|---|
| Structure | 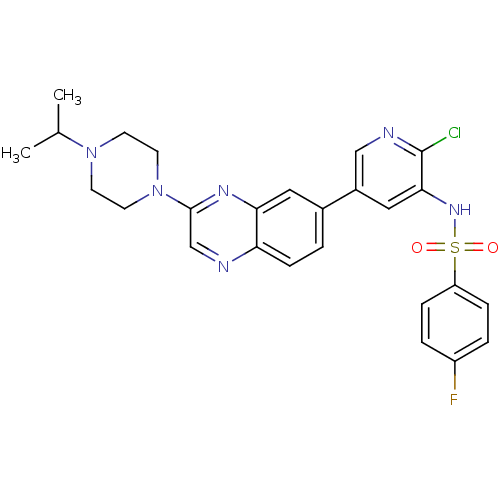
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Nishimura, N; Siegmund, A; Liu, L; Yang, K; Bryan, MC; Andrews, KL; Bo, Y; Booker, SK; Caenepeel, S; Freeman, D; Liao, H; McCarter, J; Mullady, EL; San Miguel, T; Subramanian, R; Tamayo, N; Wang, L; Whittington, DA; Zalameda, L; Zhang, N; Hughes, PE; Norman, MH Phospshoinositide 3-kinase (PI3K)/mammalian target of rapamycin (mTOR) dual inhibitors: discovery and structure-activity relationships of a series of quinoline and quinoxaline derivatives. J Med Chem54:4735-51 (2011) [PubMed] Article
Nishimura, N; Siegmund, A; Liu, L; Yang, K; Bryan, MC; Andrews, KL; Bo, Y; Booker, SK; Caenepeel, S; Freeman, D; Liao, H; McCarter, J; Mullady, EL; San Miguel, T; Subramanian, R; Tamayo, N; Wang, L; Whittington, DA; Zalameda, L; Zhang, N; Hughes, PE; Norman, MH Phospshoinositide 3-kinase (PI3K)/mammalian target of rapamycin (mTOR) dual inhibitors: discovery and structure-activity relationships of a series of quinoline and quinoxaline derivatives. J Med Chem54:4735-51 (2011) [PubMed] Article