| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Peroxisome proliferator-activated receptor delta |
|---|
| Ligand | BDBM50360740 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_796446 (CHEMBL1937761) |
|---|
| EC50 | 2700±n/a nM |
|---|
| Citation |  Nomura, M; Yumoto, K; Shinozaki, T; Isogai, S; Takano, Y; Murakami, K Discovery of cyclic amine-substituted benzoic acids as PPARa agonists. Bioorg Med Chem Lett22:334-8 (2011) [PubMed] Article Nomura, M; Yumoto, K; Shinozaki, T; Isogai, S; Takano, Y; Murakami, K Discovery of cyclic amine-substituted benzoic acids as PPARa agonists. Bioorg Med Chem Lett22:334-8 (2011) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Peroxisome proliferator-activated receptor delta |
|---|
| Name: | Peroxisome proliferator-activated receptor delta |
|---|
| Synonyms: | NR1C2 | Nuclear receptor subfamily 1 group C member 2 | PPAR-beta | PPAR-delta | PPARB | PPARD | PPARD_CANLF | Peroxisome proliferator-activated receptor beta |
|---|
| Type: | PROTEIN |
|---|
| Mol. Mass.: | 49786.42 |
|---|
| Organism: | Canis familiaris |
|---|
| Description: | ChEMBL_796446 |
|---|
| Residue: | 441 |
|---|
| Sequence: | MEQPPGEAAEVREEEEKKEVAEAEGAPELNGGPERSLPSSSYTDLSRSSSPPSLLDQLQM
GGDGASCGSLNMECRVCGDKASGFHYGVHACEGCKGFFRRTIRMKLEYEKCERICKIQKK
NRNKCQYCRFQKCVALGMSHNAIRFGRMPEAEKRKLVAGLTANEGTQHNPQVADLKAFSK
HIYNAYLKNFNMTKKKARGILTGKASHTAPFVIHDIETLWQAEKGLVWKQLVNGLPPYKE
ISVHVFYRCQCTTVETVRELTEFAKSIPSFSNLFLNDQVTLLKYGVHEAIFAMLASIVNK
DGLLVANGTGFVTREFLRSLRKPFSDIIEPKFEFAVKFNALELDDSDLALFIAAIILCGD
RPGLINVPQVEAIQDTILRALEFHLQANHPYAQYLFPKLLQKMADLRQLVTEHAQMMQRI
KKTETETSLHPLLQEIYKDMY
|
|
|
|---|
| BDBM50360740 |
|---|
| n/a |
|---|
| Name | BDBM50360740 |
|---|
| Synonyms: | CHEMBL1934476 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C23H22ClN3O3S |
|---|
| Mol. Mass. | 455.957 |
|---|
| SMILES | Cc1nc(sc1C(=O)N[C@H]1CCCN(C1)c1cccc(c1)C(O)=O)-c1ccc(Cl)cc1 |r| |
|---|
| Structure | 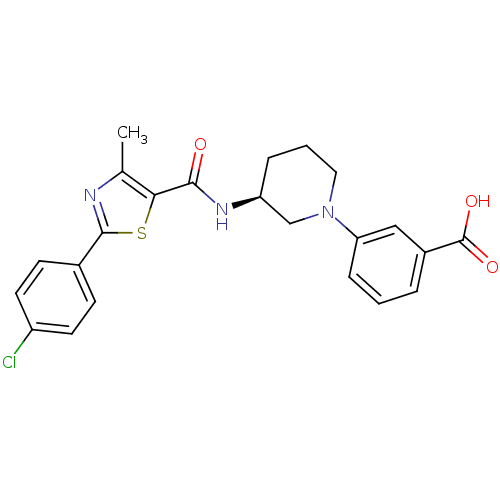
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Nomura, M; Yumoto, K; Shinozaki, T; Isogai, S; Takano, Y; Murakami, K Discovery of cyclic amine-substituted benzoic acids as PPARa agonists. Bioorg Med Chem Lett22:334-8 (2011) [PubMed] Article
Nomura, M; Yumoto, K; Shinozaki, T; Isogai, S; Takano, Y; Murakami, K Discovery of cyclic amine-substituted benzoic acids as PPARa agonists. Bioorg Med Chem Lett22:334-8 (2011) [PubMed] Article