| Reaction Details |
|---|
 | Report a problem with these data |
| Target | cAMP-dependent protein kinase catalytic subunit alpha |
|---|
| Ligand | BDBM2579 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEBML_161880 |
|---|
| IC50 | 27±n/a nM |
|---|
| Citation |  Shearer, BG; Sullivan, JP; Carter, JP; Mathew, RM; Waid, P; Connor, JR; Patch, RJ; Burch, RM Substituted 2-(aminomethyl)piperidines: a novel class of selective protein kinase C inhibitors. J Med Chem34:2928-31 (1991) [PubMed] Shearer, BG; Sullivan, JP; Carter, JP; Mathew, RM; Waid, P; Connor, JR; Patch, RJ; Burch, RM Substituted 2-(aminomethyl)piperidines: a novel class of selective protein kinase C inhibitors. J Med Chem34:2928-31 (1991) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| cAMP-dependent protein kinase catalytic subunit alpha |
|---|
| Name: | cAMP-dependent protein kinase catalytic subunit alpha |
|---|
| Synonyms: | KAPCA_RAT | PKA, catalytic subunit | Prkaca | cAMP-Dependent Protein Kinase (PKA) | cAMP-dependent protein kinase alpha-catalytic subunit |
|---|
| Type: | Enzyme catalytic subunit |
|---|
| Mol. Mass.: | 40627.70 |
|---|
| Organism: | Rattus norvegicus (rat) |
|---|
| Description: | The catalytic subunit of cAMP-dependent protein kinase (PKA) was used in the assay. It did not require cAMP for activity. |
|---|
| Residue: | 351 |
|---|
| Sequence: | MGNAAAAKKGSEQESVKEFLAKAKEDFLKKWEDPSQNTAQLDHFDRIKTLGTGSFGRVML
VKHKESGNHYAMKILDKQKVVKLKQIEHTLNEKRILQAVNFPFLVKLEFSFKDNSNLYMV
MEYVPGGEMFSHLRRIGRFSEPHARFYAAQIVLTFEYLHSLDLIYRDLKPENLLIDQQGY
IQVTDFGFAKRVKGRTWTLCGTPEYLAPEIILSKGYNKAVDWWALGVLIYEMAAGYPPFF
ADQPIQIYEKIVSGKVRFPSHFSSDLKDLLRNLLQVDLTKRFGNLKNGVNDIKNHKWFAT
TDWIAIYQRKVEAPFIPKFKGPGDTSNFDDYEEEEIRVSINEKCGKEFTEF
|
|
|
|---|
| BDBM2579 |
|---|
| n/a |
|---|
| Name | BDBM2579 |
|---|
| Synonyms: | (2S,3R,4R,6R)-3-methoxy-2-methyl-4-(methylamino)-29-oxa-1,7,17-triazaoctacyclo[12.12.2.1^{2,6}.0^{7,28}.0^{8,13}.0^{15,19}.0^{20,27}.0^{21,26}]nonacosa-8(13),9,11,14(28),15(19),20(27),21(26),22,24-nonaen-16-one | CHEMBL388978 | Staurosporin, 4 | Staurosporine | Staurosporine, 8 | US20240002365, Compound staurosporine | US9206188, Staurosporine | US9226923, Staurosporine |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C28H26N4O3 |
|---|
| Mol. Mass. | 466.531 |
|---|
| SMILES | CN[C@@H]1C[C@H]2O[C@@](C)([C@@H]1OC)n1c3ccccc3c3c4CNC(=O)c4c4c5ccccc5n2c4c13 |r| |
|---|
| Structure | 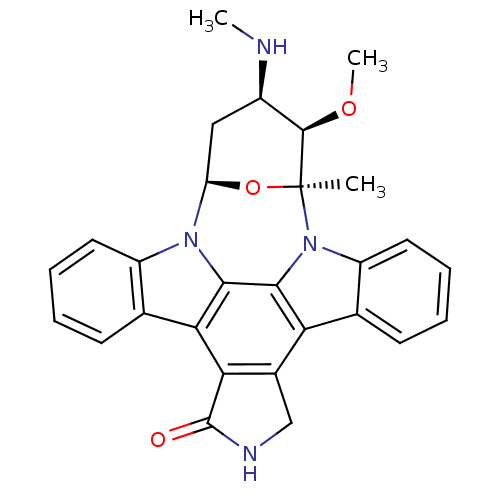
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Shearer, BG; Sullivan, JP; Carter, JP; Mathew, RM; Waid, P; Connor, JR; Patch, RJ; Burch, RM Substituted 2-(aminomethyl)piperidines: a novel class of selective protein kinase C inhibitors. J Med Chem34:2928-31 (1991) [PubMed]
Shearer, BG; Sullivan, JP; Carter, JP; Mathew, RM; Waid, P; Connor, JR; Patch, RJ; Burch, RM Substituted 2-(aminomethyl)piperidines: a novel class of selective protein kinase C inhibitors. J Med Chem34:2928-31 (1991) [PubMed]