| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Peroxisome proliferator-activated receptor gamma |
|---|
| Ligand | BDBM50175504 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_320735 (CHEMBL881208) |
|---|
| Kd | 9800±n/a nM |
|---|
| Citation |  Morphy, R; Rankovic, Z Designed multiple ligands. An emerging drug discovery paradigm. J Med Chem48:6523-43 (2005) [PubMed] Article Morphy, R; Rankovic, Z Designed multiple ligands. An emerging drug discovery paradigm. J Med Chem48:6523-43 (2005) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Peroxisome proliferator-activated receptor gamma |
|---|
| Name: | Peroxisome proliferator-activated receptor gamma |
|---|
| Synonyms: | NR1C3 | Nuclear receptor subfamily 1 group C member 3 | PPAR-gamma | PPARG | PPARG_HUMAN | Peroxisome proliferator-activated receptor | Peroxisome proliferator-activated receptor gamma (PPAR gamma) | Peroxisome proliferator-activated receptor gamma (PPARG) | Peroxisome proliferator-activated receptor gamma (PPARγ) | Peroxisome proliferator-activated receptor gamma/Nuclear receptor corepressor 2 | peroxisome proliferator-activated receptor gamma isoform 2 |
|---|
| Type: | Nuclear Receptor |
|---|
| Mol. Mass.: | 57613.46 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P37231 |
|---|
| Residue: | 505 |
|---|
| Sequence: | MGETLGDSPIDPESDSFTDTLSANISQEMTMVDTEMPFWPTNFGISSVDLSVMEDHSHSF
DIKPFTTVDFSSISTPHYEDIPFTRTDPVVADYKYDLKLQEYQSAIKVEPASPPYYSEKT
QLYNKPHEEPSNSLMAIECRVCGDKASGFHYGVHACEGCKGFFRRTIRLKLIYDRCDLNC
RIHKKSRNKCQYCRFQKCLAVGMSHNAIRFGRMPQAEKEKLLAEISSDIDQLNPESADLR
ALAKHLYDSYIKSFPLTKAKARAILTGKTTDKSPFVIYDMNSLMMGEDKIKFKHITPLQE
QSKEVAIRIFQGCQFRSVEAVQEITEYAKSIPGFVNLDLNDQVTLLKYGVHEIIYTMLAS
LMNKDGVLISEGQGFMTREFLKSLRKPFGDFMEPKFEFAVKFNALELDDSDLAIFIAVII
LSGDRPGLLNVKPIEDIQDNLLQALELQLKLNHPESSQLFAKLLQKMTDLRQIVTEHVQL
LQVIKKTETDMSLHPLLQEIYKDLY
|
|
|
|---|
| BDBM50175504 |
|---|
| n/a |
|---|
| Name | BDBM50175504 |
|---|
| Synonyms: | 3-{4-[2-(Dibenzo[b,f][1,4]thiazepin-11-yl-methyl-amino)-ethoxy]-phenyl}-2-ethoxy-propionic acid | CHEMBL194411 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C27H28N2O4S |
|---|
| Mol. Mass. | 476.587 |
|---|
| SMILES | CCOC(Cc1ccc(OCCN(C)C2=Nc3ccccc3Sc3ccccc23)cc1)C(O)=O |t:14| |
|---|
| Structure | 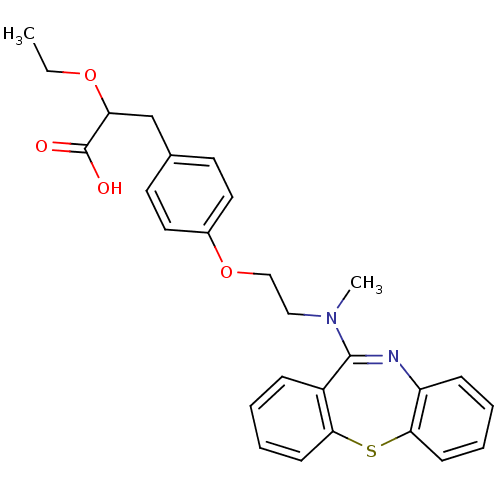
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Morphy, R; Rankovic, Z Designed multiple ligands. An emerging drug discovery paradigm. J Med Chem48:6523-43 (2005) [PubMed] Article
Morphy, R; Rankovic, Z Designed multiple ligands. An emerging drug discovery paradigm. J Med Chem48:6523-43 (2005) [PubMed] Article