| Reaction Details |
|---|
 | Report a problem with these data |
| Target | cAMP-specific 3',5'-cyclic phosphodiesterase 4C |
|---|
| Ligand | BDBM14775 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_423729 (CHEMBL855488) |
|---|
| IC50 | 14±n/a nM |
|---|
| Citation |  Duplantier, AJ; Bachert, EL; Cheng, JB; Cohan, VL; Jenkinson, TH; Kraus, KG; McKechney, MW; Pillar, JD; Watson, JW SAR of a series of 5,6-dihydro-(9H)-pyrazolo[3,4-c]-1,2,4-triazolo[4,3-alpha]pyridines as potent inhibitors of human eosinophil phosphodiesterase. J Med Chem50:344-9 (2007) [PubMed] Article Duplantier, AJ; Bachert, EL; Cheng, JB; Cohan, VL; Jenkinson, TH; Kraus, KG; McKechney, MW; Pillar, JD; Watson, JW SAR of a series of 5,6-dihydro-(9H)-pyrazolo[3,4-c]-1,2,4-triazolo[4,3-alpha]pyridines as potent inhibitors of human eosinophil phosphodiesterase. J Med Chem50:344-9 (2007) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| cAMP-specific 3',5'-cyclic phosphodiesterase 4C |
|---|
| Name: | cAMP-specific 3',5'-cyclic phosphodiesterase 4C |
|---|
| Synonyms: | 3',5'-cyclic phosphodiesterase | DPDE1 | NP_000914.2 | PDE21 | PDE4C | PDE4C1 |
|---|
| Type: | Enzyme Catalytic Domain |
|---|
| Mol. Mass.: | 79875.54 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q08493 |
|---|
| Residue: | 712 |
|---|
| Sequence: | MENLGVGEGAEACSRLSRSRGRHSMTRAPKHLWRQPRRPIRIQQRFYSDPDKSAGCRERD
LSPRPELRKSRLSWPVSSCRRFDLENGLSCGRRALDPQSSPGLGRIMQAPVPHSQRRESF
LYRSDSDYELSPKAMSRNSSVASDLHGEDMIVTPFAQVLASLRTVRSNVAALARQQCLGA
AKQGPVGNPSSSNQLPPAEDTGQKLALETLDELDWCLDQLETLQTRHSVGEMASNKFKRI
LNRELTHLSETSRSGNQVSEYISRTFLDQQTEVELPKVTAEEAPQPMSRISGLHGLCHSA
SLSSATVPRFGVQTDQEEQLAKELEDTNKWGLDVFKVAELSGNRPLTAIIFSIFQERDLL
KTFQIPADTLATYLLMLEGHYHANVAYHNSLHAADVAQSTHVLLATPALEAVFTDLEILA
ALFASAIHDVDHPGVSNQFLINTNSELALMYNDASVLENHHLAVGFKLLQAENCDIFQNL
SAKQRLSLRRMVIDMVLATDMSKHMNLLADLKTMVETKKVTSLGVLLLDNYSDRIQVLQN
LVHCADLSNPTKPLPLYRQWTDRIMAEFFQQGDRERESGLDISPMCDKHTASVEKSQVGF
IDYIAHPLWETWADLVHPDAQDLLDTLEDNREWYQSKIPRSPSDLTNPERDGPDRFQFEL
TLEEAEEEDEEEEEEGEETALAKEALELPDTELLSPEAGPDPGDLPLDNQRT
|
|
|
|---|
| BDBM14775 |
|---|
| n/a |
|---|
| Name | BDBM14775 |
|---|
| Synonyms: | 3-(cyclopentyloxy)-N-(3,5-dichloropyridin-4-yl)-4-methoxybenzamide | 3-cyclopentyloxy-N-(3,5-dichloropyridin-4-yl)-4-methoxy-benzamide | CHEMBL42126 | Cpodpmb | Piclamilast | RPR-73401 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C18H18Cl2N2O3 |
|---|
| Mol. Mass. | 381.253 |
|---|
| SMILES | COc1ccc(cc1OC1CCCC1)C(=O)Nc1c(Cl)cncc1Cl |
|---|
| Structure | 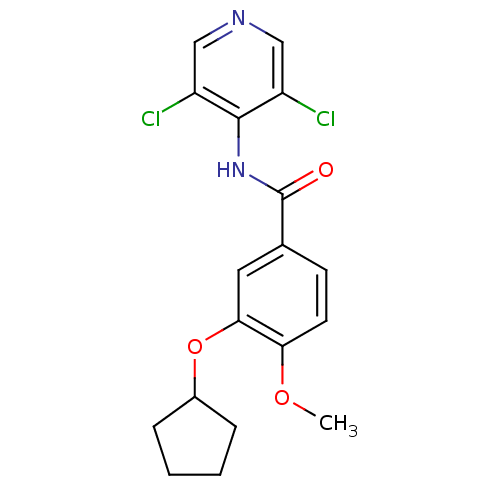
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Duplantier, AJ; Bachert, EL; Cheng, JB; Cohan, VL; Jenkinson, TH; Kraus, KG; McKechney, MW; Pillar, JD; Watson, JW SAR of a series of 5,6-dihydro-(9H)-pyrazolo[3,4-c]-1,2,4-triazolo[4,3-alpha]pyridines as potent inhibitors of human eosinophil phosphodiesterase. J Med Chem50:344-9 (2007) [PubMed] Article
Duplantier, AJ; Bachert, EL; Cheng, JB; Cohan, VL; Jenkinson, TH; Kraus, KG; McKechney, MW; Pillar, JD; Watson, JW SAR of a series of 5,6-dihydro-(9H)-pyrazolo[3,4-c]-1,2,4-triazolo[4,3-alpha]pyridines as potent inhibitors of human eosinophil phosphodiesterase. J Med Chem50:344-9 (2007) [PubMed] Article