| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Insulin-like growth factor 1 receptor |
|---|
| Ligand | BDBM50265846 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_540380 (CHEMBL1025723) |
|---|
| IC50 | 52±n/a nM |
|---|
| Citation |  Chamberlain, SD; Redman, AM; Patnaik, S; Brickhouse, K; Chew, YC; Deanda, F; Gerding, R; Lei, H; Moorthy, G; Patrick, M; Stevens, KL; Wilson, JW; Brad Shotwell, J Optimization of a series of 4,6-bis-anilino-1H-pyrrolo[2,3-d]pyrimidine inhibitors of IGF-1R: elimination of an acid-mediated decomposition pathway. Bioorg Med Chem Lett19:373-7 (2008) [PubMed] Article Chamberlain, SD; Redman, AM; Patnaik, S; Brickhouse, K; Chew, YC; Deanda, F; Gerding, R; Lei, H; Moorthy, G; Patrick, M; Stevens, KL; Wilson, JW; Brad Shotwell, J Optimization of a series of 4,6-bis-anilino-1H-pyrrolo[2,3-d]pyrimidine inhibitors of IGF-1R: elimination of an acid-mediated decomposition pathway. Bioorg Med Chem Lett19:373-7 (2008) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Insulin-like growth factor 1 receptor |
|---|
| Name: | Insulin-like growth factor 1 receptor |
|---|
| Synonyms: | CD_antigen=CD221 | IGF-I receptor | IGF1R | IGF1R_HUMAN | Insulin-like growth factor 1 receptor (IGF1R) | Insulin-like growth factor 1 receptor (IGFIR) | Insulin-like growth factor 1 receptor alpha chain | Insulin-like growth factor 1 receptor beta chain | Insulin-like growth factor I receptor | Insulin-like growth factor receptor (IGFR) |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 154776.79 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P08069 |
|---|
| Residue: | 1367 |
|---|
| Sequence: | MKSGSGGGSPTSLWGLLFLSAALSLWPTSGEICGPGIDIRNDYQQLKRLENCTVIEGYLH
ILLISKAEDYRSYRFPKLTVITEYLLLFRVAGLESLGDLFPNLTVIRGWKLFYNYALVIF
EMTNLKDIGLYNLRNITRGAIRIEKNADLCYLSTVDWSLILDAVSNNYIVGNKPPKECGD
LCPGTMEEKPMCEKTTINNEYNYRCWTTNRCQKMCPSTCGKRACTENNECCHPECLGSCS
APDNDTACVACRHYYYAGVCVPACPPNTYRFEGWRCVDRDFCANILSAESSDSEGFVIHD
GECMQECPSGFIRNGSQSMYCIPCEGPCPKVCEEEKKTKTIDSVTSAQMLQGCTIFKGNL
LINIRRGNNIASELENFMGLIEVVTGYVKIRHSHALVSLSFLKNLRLILGEEQLEGNYSF
YVLDNQNLQQLWDWDHRNLTIKAGKMYFAFNPKLCVSEIYRMEEVTGTKGRQSKGDINTR
NNGERASCESDVLHFTSTTTSKNRIIITWHRYRPPDYRDLISFTVYYKEAPFKNVTEYDG
QDACGSNSWNMVDVDLPPNKDVEPGILLHGLKPWTQYAVYVKAVTLTMVENDHIRGAKSE
ILYIRTNASVPSIPLDVLSASNSSSQLIVKWNPPSLPNGNLSYYIVRWQRQPQDGYLYRH
NYCSKDKIPIRKYADGTIDIEEVTENPKTEVCGGEKGPCCACPKTEAEKQAEKEEAEYRK
VFENFLHNSIFVPRPERKRRDVMQVANTTMSSRSRNTTAADTYNITDPEELETEYPFFES
RVDNKERTVISNLRPFTLYRIDIHSCNHEAEKLGCSASNFVFARTMPAEGADDIPGPVTW
EPRPENSIFLKWPEPENPNGLILMYEIKYGSQVEDQRECVSRQEYRKYGGAKLNRLNPGN
YTARIQATSLSGNGSWTDPVFFYVQAKTGYENFIHLIIALPVAVLLIVGGLVIMLYVFHR
KRNNSRLGNGVLYASVNPEYFSAADVYVPDEWEVAREKITMSRELGQGSFGMVYEGVAKG
VVKDEPETRVAIKTVNEAASMRERIEFLNEASVMKEFNCHHVVRLLGVVSQGQPTLVIME
LMTRGDLKSYLRSLRPEMENNPVLAPPSLSKMIQMAGEIADGMAYLNANKFVHRDLAARN
CMVAEDFTVKIGDFGMTRDIYETDYYRKGGKGLLPVRWMSPESLKDGVFTTYSDVWSFGV
VLWEIATLAEQPYQGLSNEQVLRFVMEGGLLDKPDNCPDMLFELMRMCWQYNPKMRPSFL
EIISSIKEEMEPGFREVSFYYSEENKLPEPEELDLEPENMESVPLDPSASSSSLPLPDRH
SGHKAENGPGPGVLVLRASFDERQPYAHMNGGRKNERALPLPQSSTC
|
|
|
|---|
| BDBM50265846 |
|---|
| n/a |
|---|
| Name | BDBM50265846 |
|---|
| Synonyms: | 3-(2-(2-methoxy-4-(1-propylpiperidin-4-yl)phenylamino)-7H-pyrrolo[2,3-d]pyrimidin-4-ylamino)thiophene-2-carboxamide | CHEMBL515846 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C26H31N7O2S |
|---|
| Mol. Mass. | 505.635 |
|---|
| SMILES | CCCN1CCC(CC1)c1ccc(Nc2nc(Nc3ccsc3C(N)=O)c3cc[nH]c3n2)c(OC)c1 |
|---|
| Structure | 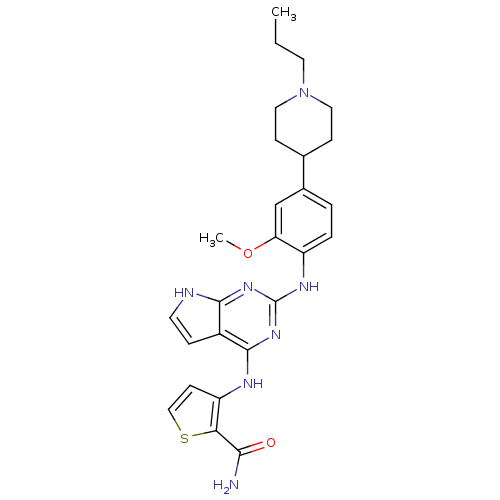
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Chamberlain, SD; Redman, AM; Patnaik, S; Brickhouse, K; Chew, YC; Deanda, F; Gerding, R; Lei, H; Moorthy, G; Patrick, M; Stevens, KL; Wilson, JW; Brad Shotwell, J Optimization of a series of 4,6-bis-anilino-1H-pyrrolo[2,3-d]pyrimidine inhibitors of IGF-1R: elimination of an acid-mediated decomposition pathway. Bioorg Med Chem Lett19:373-7 (2008) [PubMed] Article
Chamberlain, SD; Redman, AM; Patnaik, S; Brickhouse, K; Chew, YC; Deanda, F; Gerding, R; Lei, H; Moorthy, G; Patrick, M; Stevens, KL; Wilson, JW; Brad Shotwell, J Optimization of a series of 4,6-bis-anilino-1H-pyrrolo[2,3-d]pyrimidine inhibitors of IGF-1R: elimination of an acid-mediated decomposition pathway. Bioorg Med Chem Lett19:373-7 (2008) [PubMed] Article