| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Cyclin-dependent kinase 9 |
|---|
| Ligand | BDBM4851 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_586445 (CHEMBL1063701) |
|---|
| Kd | >10000±n/a nM |
|---|
| Citation |  Karaman, MW; Herrgard, S; Treiber, DK; Gallant, P; Atteridge, CE; Campbell, BT; Chan, KW; Ciceri, P; Davis, MI; Edeen, PT; Faraoni, R; Floyd, M; Hunt, JP; Lockhart, DJ; Milanov, ZV; Morrison, MJ; Pallares, G; Patel, HK; Pritchard, S; Wodicka, LM; Zarrinkar, PP A quantitative analysis of kinase inhibitor selectivity. Nat Biotechnol26:127-32 (2008) [PubMed] Article Karaman, MW; Herrgard, S; Treiber, DK; Gallant, P; Atteridge, CE; Campbell, BT; Chan, KW; Ciceri, P; Davis, MI; Edeen, PT; Faraoni, R; Floyd, M; Hunt, JP; Lockhart, DJ; Milanov, ZV; Morrison, MJ; Pallares, G; Patel, HK; Pritchard, S; Wodicka, LM; Zarrinkar, PP A quantitative analysis of kinase inhibitor selectivity. Nat Biotechnol26:127-32 (2008) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Cyclin-dependent kinase 9 |
|---|
| Name: | Cyclin-dependent kinase 9 |
|---|
| Synonyms: | C-2K | CDC2L4 | CDK9 | CDK9_HUMAN | Cell division cycle 2-like protein kinase 4 | Cell division protein kinase 9 | Cyclin-Dependent Kinase 9 (CDK9) | Cyclin-dependent kinase 9 (CDK9) | Serine/threonine-protein kinase PITALRE | TAK | Tat-associated kinase complex catalytic subunit |
|---|
| Type: | Enzyme Subunit |
|---|
| Mol. Mass.: | 42789.13 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | n/a |
|---|
| Residue: | 372 |
|---|
| Sequence: | MAKQYDSVECPFCDEVSKYEKLAKIGQGTFGEVFKARHRKTGQKVALKKVLMENEKEGFP
ITALREIKILQLLKHENVVNLIEICRTKASPYNRCKGSIYLVFDFCEHDLAGLLSNVLVK
FTLSEIKRVMQMLLNGLYYIHRNKILHRDMKAANVLITRDGVLKLADFGLARAFSLAKNS
QPNRYTNRVVTLWYRPPELLLGERDYGPPIDLWGAGCIMAEMWTRSPIMQGNTEQHQLAL
ISQLCGSITPEVWPNVDNYELYEKLELVKGQKRKVKDRLKAYVRDPYALDLIDKLLVLDP
AQRIDSDDALNHDFFWSDPMPSDLKGMLSTHLTSMFEYLAPPRRKGSQITQQSTNQSRNP
ATTNQTEFERVF
|
|
|
|---|
| BDBM4851 |
|---|
| n/a |
|---|
| Name | BDBM4851 |
|---|
| Synonyms: | (4-chlorophenyl)-[4-(4-pyridylmethyl)phthalazin-1-yl]amine | (A) PTK787 | CGP 79787 | CHEMBL101253 | CHEMBL75232 | N-(4-chlorophenyl)-4-(pyridin-4-ylmethyl)-1-phthalazinamine | N-(4-chlorophenyl)-4-(pyridin-4-ylmethyl)phthalazin-1-amine | PTK-787 | PTK787 | ZK222584 | cid_151194 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C20H15ClN4 |
|---|
| Mol. Mass. | 346.813 |
|---|
| SMILES | Clc1ccc(Nc2nnc(Cc3ccncc3)c3ccccc23)cc1 |
|---|
| Structure | 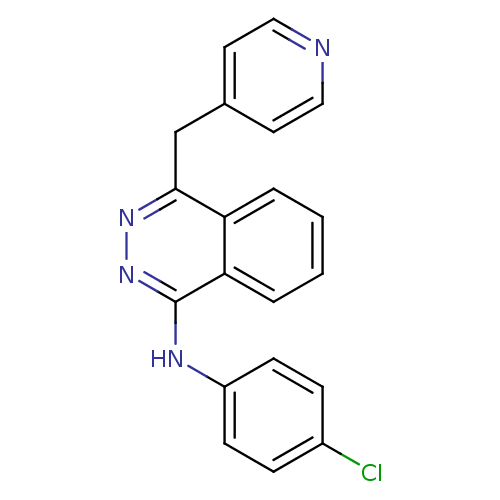
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Karaman, MW; Herrgard, S; Treiber, DK; Gallant, P; Atteridge, CE; Campbell, BT; Chan, KW; Ciceri, P; Davis, MI; Edeen, PT; Faraoni, R; Floyd, M; Hunt, JP; Lockhart, DJ; Milanov, ZV; Morrison, MJ; Pallares, G; Patel, HK; Pritchard, S; Wodicka, LM; Zarrinkar, PP A quantitative analysis of kinase inhibitor selectivity. Nat Biotechnol26:127-32 (2008) [PubMed] Article
Karaman, MW; Herrgard, S; Treiber, DK; Gallant, P; Atteridge, CE; Campbell, BT; Chan, KW; Ciceri, P; Davis, MI; Edeen, PT; Faraoni, R; Floyd, M; Hunt, JP; Lockhart, DJ; Milanov, ZV; Morrison, MJ; Pallares, G; Patel, HK; Pritchard, S; Wodicka, LM; Zarrinkar, PP A quantitative analysis of kinase inhibitor selectivity. Nat Biotechnol26:127-32 (2008) [PubMed] Article