

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Cannabinoid receptor 1 | ||
| Ligand | BDBM50298953 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_588141 (CHEMBL1049258) | ||
| Ki | 117±n/a nM | ||
| Citation |  Silvestri, R; Ligresti, A; La Regina, G; Piscitelli, F; Gatti, V; Brizzi, A; Pasquini, S; Lavecchia, A; Allarà, M; Fantini, N; Carai, MA; Novellino, E; Colombo, G; Di Marzo, V; Corelli, F Synthesis, cannabinoid receptor affinity, molecular modeling studies and in vivo pharmacological evaluation of new substituted 1-aryl-5-(1H-pyrrol-1-yl)-1H-pyrazole-3-carboxamides. 2. Effect of the 3-carboxamide substituent on the affinity and selectivity profile. Bioorg Med Chem17:5549-64 (2009) [PubMed] Article Silvestri, R; Ligresti, A; La Regina, G; Piscitelli, F; Gatti, V; Brizzi, A; Pasquini, S; Lavecchia, A; Allarà, M; Fantini, N; Carai, MA; Novellino, E; Colombo, G; Di Marzo, V; Corelli, F Synthesis, cannabinoid receptor affinity, molecular modeling studies and in vivo pharmacological evaluation of new substituted 1-aryl-5-(1H-pyrrol-1-yl)-1H-pyrazole-3-carboxamides. 2. Effect of the 3-carboxamide substituent on the affinity and selectivity profile. Bioorg Med Chem17:5549-64 (2009) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Cannabinoid receptor 1 | |||
| Name: | Cannabinoid receptor 1 | ||
| Synonyms: | CANN6 | CANNABINOID CB1 | CB-R | CB1 | CNR | CNR1 | CNR1_HUMAN | Cannabinoid CB1 receptor | Cannabinoid receptor | Cannabinoid receptor 1 (CB1) | Cannabinoid receptor 1 (brain) | ||
| Type: | G Protein-Coupled Receptor (GPCR) | ||
| Mol. Mass.: | 52868.96 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | P21554 | ||
| Residue: | 472 | ||
| Sequence: |
| ||
| BDBM50298953 | |||
| n/a | |||
| Name | BDBM50298953 | ||
| Synonyms: | CHEMBL574032 | N-[2-(3,4-Dichlorophenyl)ethyl]1-(2,4-dichlorophenyl)-5-(2,5-dimethyl-1H-pyrrol-1-yl)-4-methyl-1H-pyrazole-3-carboxamide | ||
| Type | Small organic molecule | ||
| Emp. Form. | C25H22Cl4N4O | ||
| Mol. Mass. | 536.28 | ||
| SMILES | Cc1ccc(C)n1-c1c(C)c(nn1-c1ccc(Cl)cc1Cl)C(=O)NCCc1ccc(Cl)c(Cl)c1 |(4.67,-7.31,;5.6,-6.07,;7.14,-6.08,;7.62,-4.62,;6.38,-3.7,;6.39,-2.16,;5.13,-4.6,;3.67,-4.11,;3.2,-2.64,;4.11,-1.4,;1.65,-2.62,;1.17,-4.09,;2.41,-5.01,;2.4,-6.55,;3.73,-7.34,;3.72,-8.88,;2.38,-9.64,;2.37,-11.18,;1.05,-8.86,;1.06,-7.32,;-.27,-6.53,;.76,-1.37,;1.39,.03,;-.78,-1.52,;-1.68,-.27,;-3.21,-.41,;-4.11,.84,;-3.48,2.24,;-4.38,3.49,;-5.91,3.35,;-6.8,4.6,;-6.55,1.94,;-8.08,1.79,;-5.65,.69,)| | ||
| Structure | 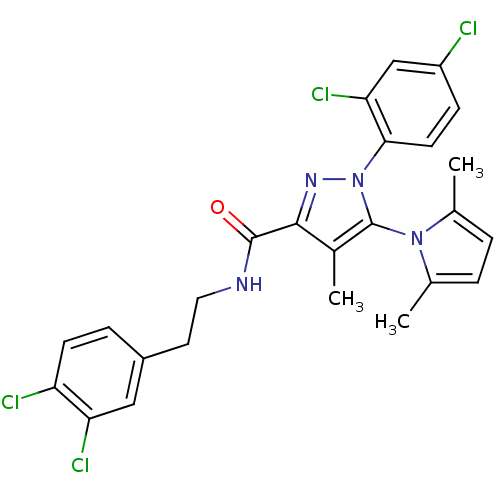
| ||