

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Muscarinic acetylcholine receptor M3 | ||
| Ligand | BDBM50381646 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_815844 (CHEMBL2026107) | ||
| Kd | 0.0501±n/a nM | ||
| Citation |  Lainé, DI; Yan, H; Xie, H; Davis, RS; Dufour, J; Widdowson, KL; Palovich, MR; Wan, Z; Foley, JJ; Schmidt, DB; Hunsberger, GE; Burman, M; Bacon, AM; Webb, EF; Luttmann, MA; Salmon, M; Sarau, HM; Umbrecht, ST; Landis, PS; Peck, BJ; Busch-Petersen, J Design, synthesis and structure-activity relationship of N-substituted tropane muscarinic acetylcholine receptor antagonists. Bioorg Med Chem Lett22:3366-9 (2012) [PubMed] Article Lainé, DI; Yan, H; Xie, H; Davis, RS; Dufour, J; Widdowson, KL; Palovich, MR; Wan, Z; Foley, JJ; Schmidt, DB; Hunsberger, GE; Burman, M; Bacon, AM; Webb, EF; Luttmann, MA; Salmon, M; Sarau, HM; Umbrecht, ST; Landis, PS; Peck, BJ; Busch-Petersen, J Design, synthesis and structure-activity relationship of N-substituted tropane muscarinic acetylcholine receptor antagonists. Bioorg Med Chem Lett22:3366-9 (2012) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Muscarinic acetylcholine receptor M3 | |||
| Name: | Muscarinic acetylcholine receptor M3 | ||
| Synonyms: | ACM3_HUMAN | CHRM3 | Cholinergic, muscarinic M3 | Muscarinic Receptors M3 | Muscarinic receptor M3 | RecName: Full=Muscarinic acetylcholine receptor M3 | ||
| Type: | Enzyme | ||
| Mol. Mass.: | 66151.03 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | P20309 | ||
| Residue: | 590 | ||
| Sequence: |
| ||
| BDBM50381646 | |||
| n/a | |||
| Name | BDBM50381646 | ||
| Synonyms: | CHEMBL2023749 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C27H33N2 | ||
| Mol. Mass. | 385.5638 | ||
| SMILES | C[N+]1(CC2CC2)C2CCC1C[C@@H](CC(C#N)(c1ccccc1)c1ccccc1)C2 |r,wD:11.13,TLB:0:1:11.10.28:8.7,THB:2:1:11.10.28:8.7,12:11:1:8.7,(5.68,2.81,;7.18,2.4,;7.94,3.74,;7.16,5.07,;7.16,6.61,;5.83,5.83,;8.23,1.57,;7.63,.23,;6.45,-.48,;7.44,.86,;9.28,.86,;10.26,.06,;11.04,-1.28,;12.58,-1.27,;12.97,.23,;13.36,1.72,;13.35,-2.61,;12.58,-3.94,;13.35,-5.27,;14.9,-5.27,;15.66,-3.92,;14.89,-2.59,;14.06,-.86,;15.15,-1.95,;16.63,-1.54,;17.02,-.05,;15.92,1.03,;14.44,.62,;10,1.6,)| | ||
| Structure | 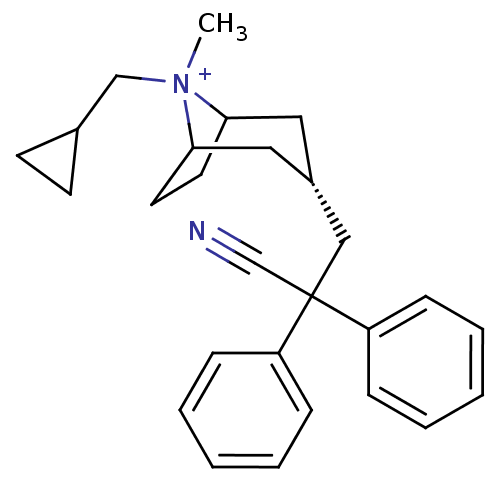
| ||