| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Fatty-acid amide hydrolase 1 [30-579] |
|---|
| Ligand | BDBM50402675 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_886874 (CHEMBL2211372) |
|---|
| IC50 | 0.7±n/a nM |
|---|
| Citation |  Tichenor, MS; Keith, JM; Jones, WM; Pierce, JM; Merit, J; Hawryluk, N; Seierstad, M; Palmer, JA; Webb, M; Karbarz, MJ; Wilson, SJ; Wennerholm, ML; Woestenborghs, F; Beerens, D; Luo, L; Brown, SM; Boeck, MD; Chaplan, SR; Breitenbucher, JG Heteroaryl urea inhibitors of fatty acid amide hydrolase: structure-mutagenicity relationships for arylamine metabolites. Bioorg Med Chem Lett22:7357-62 (2012) [PubMed] Article Tichenor, MS; Keith, JM; Jones, WM; Pierce, JM; Merit, J; Hawryluk, N; Seierstad, M; Palmer, JA; Webb, M; Karbarz, MJ; Wilson, SJ; Wennerholm, ML; Woestenborghs, F; Beerens, D; Luo, L; Brown, SM; Boeck, MD; Chaplan, SR; Breitenbucher, JG Heteroaryl urea inhibitors of fatty acid amide hydrolase: structure-mutagenicity relationships for arylamine metabolites. Bioorg Med Chem Lett22:7357-62 (2012) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Fatty-acid amide hydrolase 1 [30-579] |
|---|
| Name: | Fatty-acid amide hydrolase 1 [30-579] |
|---|
| Synonyms: | Anandamide amidohydrolase 1 | FAAH1_RAT | Faah | Faah1 | Fatty Acid Amide Hydrolase | Fatty Acid Amide Hydrolic, FAAH | Fatty-acid amide hydrolase (FAAH) | Fatty-acid amide hydrolase 1 | Fatty-acid amide hydrolase 1 (FAAH) | Fatty-acid amide hydrolase 1 (aa 30-579) | Oleamide hydrolase 1 |
|---|
| Type: | Single-pass membrane protein; homodimer |
|---|
| Mol. Mass.: | 60474.00 |
|---|
| Organism: | Rattus norvegicus (rat) |
|---|
| Description: | P97612 (aa 30-579) |
|---|
| Residue: | 550 |
|---|
| Sequence: | RWTGRQKARGAATRARQKQRASLETMDKAVQRFRLQNPDLDSEALLTLPLLQLVQKLQSG
ELSPEAVFFTYLGKAWEVNKGTNCVTSYLTDCETQLSQAPRQGLLYGVPVSLKECFSYKG
HDSTLGLSLNEGMPSESDCVVVQVLKLQGAVPFVHTNVPQSMLSFDCSNPLFGQTMNPWK
SSKSPGGSSGGEGALIGSGGSPLGLGTDIGGSIRFPSAFCGICGLKPTGNRLSKSGLKGC
VYGQTAVQLSLGPMARDVESLALCLKALLCEHLFTLDPTVPPLPFREEVYRSSRPLRVGY
YETDNYTMPSPAMRRALIETKQRLEAAGHTLIPFLPNNIPYALEVLSAGGLFSDGGRSFL
QNFKGDFVDPCLGDLILILRLPSWFKRLLSLLLKPLFPRLAAFLNSMRPRSAEKLWKLQH
EIEMYRQSVIAQWKAMNLDVLLTPMLGPALDLNTPGRATGAISYTVLYNCLDFPAGVVPV
TTVTAEDDAQMELYKGYFGDIWDIILKKAMKNSVGLPVAVQCVALPWQEELCLRFMREVE
QLMTPQKQPS
|
|
|
|---|
| BDBM50402675 |
|---|
| n/a |
|---|
| Name | BDBM50402675 |
|---|
| Synonyms: | CHEMBL2207348 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C24H22ClN5O3 |
|---|
| Mol. Mass. | 463.916 |
|---|
| SMILES | Clc1ccc(Oc2cccc(CN3CCN(CC3)C(=O)Nc3noc4cnccc34)c2)cc1 |
|---|
| Structure | 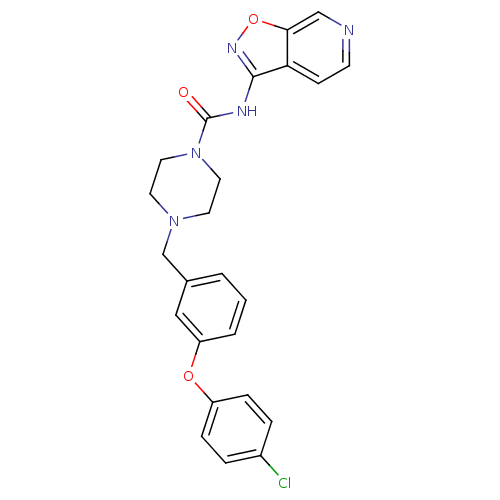
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Tichenor, MS; Keith, JM; Jones, WM; Pierce, JM; Merit, J; Hawryluk, N; Seierstad, M; Palmer, JA; Webb, M; Karbarz, MJ; Wilson, SJ; Wennerholm, ML; Woestenborghs, F; Beerens, D; Luo, L; Brown, SM; Boeck, MD; Chaplan, SR; Breitenbucher, JG Heteroaryl urea inhibitors of fatty acid amide hydrolase: structure-mutagenicity relationships for arylamine metabolites. Bioorg Med Chem Lett22:7357-62 (2012) [PubMed] Article
Tichenor, MS; Keith, JM; Jones, WM; Pierce, JM; Merit, J; Hawryluk, N; Seierstad, M; Palmer, JA; Webb, M; Karbarz, MJ; Wilson, SJ; Wennerholm, ML; Woestenborghs, F; Beerens, D; Luo, L; Brown, SM; Boeck, MD; Chaplan, SR; Breitenbucher, JG Heteroaryl urea inhibitors of fatty acid amide hydrolase: structure-mutagenicity relationships for arylamine metabolites. Bioorg Med Chem Lett22:7357-62 (2012) [PubMed] Article