| Reaction Details |
|---|
 | Report a problem with these data |
| Target | TGF-beta receptor type-1 |
|---|
| Ligand | BDBM50262079 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_961930 (CHEMBL2390521) |
|---|
| IC50 | 565±n/a nM |
|---|
| Citation |  Engers, DW; Frist, AY; Lindsley, CW; Hong, CC; Hopkins, CR Synthesis and structure-activity relationships of a novel and selective bone morphogenetic protein receptor (BMP) inhibitor derived from the pyrazolo[1.5-a]pyrimidine scaffold of dorsomorphin: the discovery of ML347 as an ALK2 versus ALK3 selective MLPCN probe. Bioorg Med Chem Lett23:3248-52 (2013) [PubMed] Article Engers, DW; Frist, AY; Lindsley, CW; Hong, CC; Hopkins, CR Synthesis and structure-activity relationships of a novel and selective bone morphogenetic protein receptor (BMP) inhibitor derived from the pyrazolo[1.5-a]pyrimidine scaffold of dorsomorphin: the discovery of ML347 as an ALK2 versus ALK3 selective MLPCN probe. Bioorg Med Chem Lett23:3248-52 (2013) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| TGF-beta receptor type-1 |
|---|
| Name: | TGF-beta receptor type-1 |
|---|
| Synonyms: | ALK-5 | ALK5 | Activin A receptor type II-like protein kinase of 53kD | Activin receptor-like kinase 5 | SKR4 | Serine/threonine-protein kinase receptor R4 | TGF-beta receptor type I | TGF-beta type I receptor | TGFBR1 | TGFR-1 | TGFR1_HUMAN | TbetaR-I | Transforming growth factor-beta receptor type I |
|---|
| Type: | enzyme |
|---|
| Mol. Mass.: | 55968.24 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P36897 |
|---|
| Residue: | 503 |
|---|
| Sequence: | MEAAVAAPRPRLLLLVLAAAAAAAAALLPGATALQCFCHLCTKDNFTCVTDGLCFVSVTE
TTDKVIHNSMCIAEIDLIPRDRPFVCAPSSKTGSVTTTYCCNQDHCNKIELPTTVKSSPG
LGPVELAAVIAGPVCFVCISLMLMVYICHNRTVIHHRVPNEEDPSLDRPFISEGTTLKDL
IYDMTTSGSGSGLPLLVQRTIARTIVLQESIGKGRFGEVWRGKWRGEEVAVKIFSSREER
SWFREAEIYQTVMLRHENILGFIAADNKDNGTWTQLWLVSDYHEHGSLFDYLNRYTVTVE
GMIKLALSTASGLAHLHMEIVGTQGKPAIAHRDLKSKNILVKKNGTCCIADLGLAVRHDS
ATDTIDIAPNHRVGTKRYMAPEVLDDSINMKHFESFKRADIYAMGLVFWEIARRCSIGGI
HEDYQLPYYDLVPSDPSVEEMRKVVCEQKLRPNIPNRWQSCEALRVMAKIMRECWYANGA
ARLTALRIKKTLSQLSQQEGIKM
|
|
|
|---|
| BDBM50262079 |
|---|
| n/a |
|---|
| Name | BDBM50262079 |
|---|
| Synonyms: | 4-(6-(4-(piperazin-1-yl)phenyl)pyrazolo[1,5-a]pyrimidin-3-yl)quinoline | 4-[6-(4-piperazin-1-ylphenyl)pyrazolo[1,5-a]pyrimidin-3-yl]quinoline | CHEMBL513147 | LDN-193189 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C25H22N6 |
|---|
| Mol. Mass. | 406.4824 |
|---|
| SMILES | C1CN(CCN1)c1ccc(cc1)-c1cnc2c(cnn2c1)-c1ccnc2ccccc12 |
|---|
| Structure | 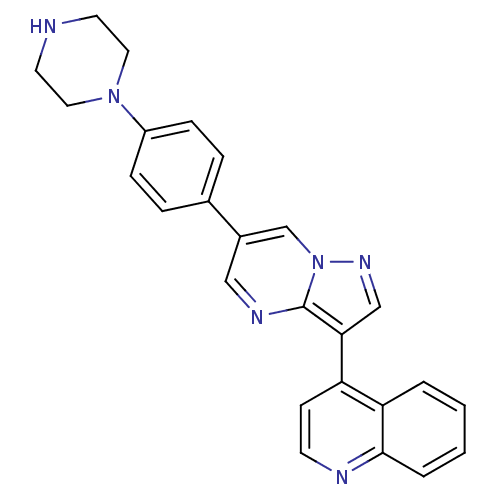
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Engers, DW; Frist, AY; Lindsley, CW; Hong, CC; Hopkins, CR Synthesis and structure-activity relationships of a novel and selective bone morphogenetic protein receptor (BMP) inhibitor derived from the pyrazolo[1.5-a]pyrimidine scaffold of dorsomorphin: the discovery of ML347 as an ALK2 versus ALK3 selective MLPCN probe. Bioorg Med Chem Lett23:3248-52 (2013) [PubMed] Article
Engers, DW; Frist, AY; Lindsley, CW; Hong, CC; Hopkins, CR Synthesis and structure-activity relationships of a novel and selective bone morphogenetic protein receptor (BMP) inhibitor derived from the pyrazolo[1.5-a]pyrimidine scaffold of dorsomorphin: the discovery of ML347 as an ALK2 versus ALK3 selective MLPCN probe. Bioorg Med Chem Lett23:3248-52 (2013) [PubMed] Article