| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Cyclin-dependent kinase/G2/mitotic-specific cyclin- 1 |
|---|
| Ligand | BDBM50440020 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_981896 (CHEMBL2427769) |
|---|
| IC50 | 1520±n/a nM |
|---|
| Citation |  Palmer, WS; Alam, M; Arzeno, HB; Chang, KC; Dunn, JP; Goldstein, DM; Gong, L; Goyal, B; Hermann, JC; Hogg, JH; Hsieh, G; Jahangir, A; Janson, C; Jin, S; Ursula Kammlott, R; Kuglstatter, A; Lukacs, C; Michoud, C; Niu, L; Reuter, DC; Shao, A; Silva, T; Trejo-Martin, TA; Stein, K; Tan, YC; Tivitmahaisoon, P; Tran, P; Wagner, P; Weller, P; Wu, SY Development of amino-pyrimidine inhibitors of c-Jun N-terminal kinase (JNK): kinase profiling guided optimization of a 1,2,3-benzotriazole lead. Bioorg Med Chem Lett23:1486-92 (2013) [PubMed] Article Palmer, WS; Alam, M; Arzeno, HB; Chang, KC; Dunn, JP; Goldstein, DM; Gong, L; Goyal, B; Hermann, JC; Hogg, JH; Hsieh, G; Jahangir, A; Janson, C; Jin, S; Ursula Kammlott, R; Kuglstatter, A; Lukacs, C; Michoud, C; Niu, L; Reuter, DC; Shao, A; Silva, T; Trejo-Martin, TA; Stein, K; Tan, YC; Tivitmahaisoon, P; Tran, P; Wagner, P; Weller, P; Wu, SY Development of amino-pyrimidine inhibitors of c-Jun N-terminal kinase (JNK): kinase profiling guided optimization of a 1,2,3-benzotriazole lead. Bioorg Med Chem Lett23:1486-92 (2013) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Cyclin-dependent kinase/G2/mitotic-specific cyclin- 1 |
|---|
| Name: | Cyclin-dependent kinase/G2/mitotic-specific cyclin- 1 |
|---|
| Synonyms: | CDK1/B | CDK1/Cyclin B | CDK1/CyclinB | Cyclin B/Dependent Kinase 1 (CDK1) | Cyclin-Dependent Kinase 1 (CDK1) | Cyclin-dependent kinase 1/cyclin B1 | Cyclin-dependent kinase/G2/mitotic-specific cyclin 1 | cdk1 |
|---|
| Type: | Protein Complex |
|---|
| Mol. Mass.: | n/a |
|---|
| Description: | CDK1/Cyclin B complexes were purified from insect cells co-infected with baculovirus vectors containing each of the components. |
|---|
| Components: | This complex has 2 components. |
|---|
| Component 1 |
| Name: | Cyclin-dependent kinase 1 |
|---|
| Synonyms: | CDC2 | CDC28A | CDK1 | CDK1_HUMAN | CDKN1 | Cell division control protein 2 homolog | Cell division protein kinase 1 | Cyclin-dependent kinase 1 (CDK1) | P34CDC2 | p34 protein kinase |
|---|
| Type: | Enzyme Subunit |
|---|
| Mol. Mass.: | 34101.08 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P06493 |
|---|
| Residue: | 297 |
|---|
| Sequence: | MEDYTKIEKIGEGTYGVVYKGRHKTTGQVVAMKKIRLESEEEGVPSTAIREISLLKELRH
PNIVSLQDVLMQDSRLYLIFEFLSMDLKKYLDSIPPGQYMDSSLVKSYLYQILQGIVFCH
SRRVLHRDLKPQNLLIDDKGTIKLADFGLARAFGIPIRVYTHEVVTLWYRSPEVLLGSAR
YSTPVDIWSIGTIFAELATKKPLFHGDSEIDQLFRIFRALGTPNNEVWPEVESLQDYKNT
FPKWKPGSLASHVKNLDENGLDLLSKMLIYDPAKRISGKMALNHPYFNDLDNQIKKM
|
|
|
|---|
| Component 2 |
| Name: | G2/mitotic-specific cyclin-B1 |
|---|
| Synonyms: | CCNB | CCNB1 | CCNB1_HUMAN |
|---|
| Type: | Enzyme Subunit |
|---|
| Mol. Mass.: | 48340.95 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P14635 |
|---|
| Residue: | 433 |
|---|
| Sequence: | MALRVTRNSKINAENKAKINMAGAKRVPTAPAATSKPGLRPRTALGDIGNKVSEQLQAKM
PMKKEAKPSATGKVIDKKLPKPLEKVPMLVPVPVSEPVPEPEPEPEPEPVKEEKLSPEPI
LVDTASPSPMETSGCAPAEEDLCQAFSDVILAVNDVDAEDGADPNLCSEYVKDIYAYLRQ
LEEEQAVRPKYLLGREVTGNMRAILIDWLVQVQMKFRLLQETMYMTVSIIDRFMQNNCVP
KKMLQLVGVTAMFIASKYEEMYPPEIGDFAFVTDNTYTKHQIRQMEMKILRALNFGLGRP
LPLHFLRRASKIGEVDVEQHTLAKYLMELTMLDYDMVHFPPSQIAAGAFCLALKILDNGE
WTPTLQHYLSYTEESLLPVMQHLAKNVVMVNQGLTKHMTVKNKYATSKHAKISTLPQLNS
ALVQDLAKAVAKV
|
|
|
|---|
| BDBM50440020 |
|---|
| n/a |
|---|
| Name | BDBM50440020 |
|---|
| Synonyms: | CHEMBL2425655 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C17H21N7O2S |
|---|
| Mol. Mass. | 387.459 |
|---|
| SMILES | CS(=O)(=O)N[C@H]1CC[C@@H](CC1)Nc1nccc(n1)-n1nnc2ccccc12 |r,wU:8.11,wD:5.4,(4.67,-9.66,;3.88,-8.32,;3.09,-6.98,;2.54,-9.1,;5.22,-7.53,;6.57,-8.3,;6.58,-9.85,;7.92,-10.61,;9.25,-9.83,;9.24,-8.29,;7.92,-7.53,;10.6,-10.6,;11.94,-9.82,;11.94,-8.28,;13.27,-7.51,;14.6,-8.27,;14.6,-9.82,;13.28,-10.59,;15.94,-10.58,;17.36,-10.03,;18.33,-11.22,;17.5,-12.52,;17.91,-14,;16.83,-15.08,;15.33,-14.67,;14.94,-13.22,;16.02,-12.12,)| |
|---|
| Structure | 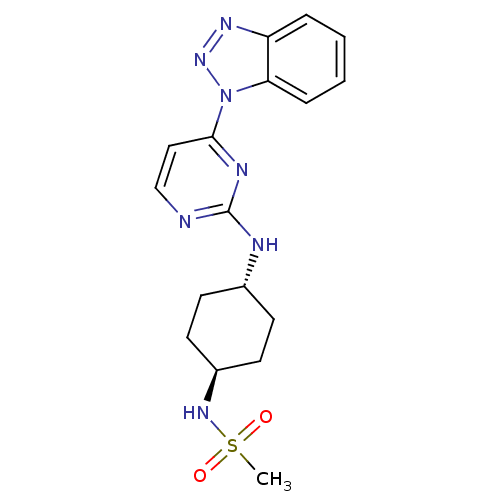
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Palmer, WS; Alam, M; Arzeno, HB; Chang, KC; Dunn, JP; Goldstein, DM; Gong, L; Goyal, B; Hermann, JC; Hogg, JH; Hsieh, G; Jahangir, A; Janson, C; Jin, S; Ursula Kammlott, R; Kuglstatter, A; Lukacs, C; Michoud, C; Niu, L; Reuter, DC; Shao, A; Silva, T; Trejo-Martin, TA; Stein, K; Tan, YC; Tivitmahaisoon, P; Tran, P; Wagner, P; Weller, P; Wu, SY Development of amino-pyrimidine inhibitors of c-Jun N-terminal kinase (JNK): kinase profiling guided optimization of a 1,2,3-benzotriazole lead. Bioorg Med Chem Lett23:1486-92 (2013) [PubMed] Article
Palmer, WS; Alam, M; Arzeno, HB; Chang, KC; Dunn, JP; Goldstein, DM; Gong, L; Goyal, B; Hermann, JC; Hogg, JH; Hsieh, G; Jahangir, A; Janson, C; Jin, S; Ursula Kammlott, R; Kuglstatter, A; Lukacs, C; Michoud, C; Niu, L; Reuter, DC; Shao, A; Silva, T; Trejo-Martin, TA; Stein, K; Tan, YC; Tivitmahaisoon, P; Tran, P; Wagner, P; Weller, P; Wu, SY Development of amino-pyrimidine inhibitors of c-Jun N-terminal kinase (JNK): kinase profiling guided optimization of a 1,2,3-benzotriazole lead. Bioorg Med Chem Lett23:1486-92 (2013) [PubMed] Article