

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Induced myeloid leukemia cell differentiation protein Mcl-1 | ||
| Ligand | BDBM50078165 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_1472596 (CHEMBL3417961) | ||
| Ki | 98±n/a nM | ||
| Citation |  Bruncko, M; Wang, L; Sheppard, GS; Phillips, DC; Tahir, SK; Xue, J; Erickson, S; Fidanze, S; Fry, E; Hasvold, L; Jenkins, GJ; Jin, S; Judge, RA; Kovar, PJ; Madar, D; Nimmer, P; Park, C; Petros, AM; Rosenberg, SH; Smith, ML; Song, X; Sun, C; Tao, ZF; Wang, X; Xiao, Y; Zhang, H; Tse, C; Leverson, JD; Elmore, SW; Souers, AJ Structure-guided design of a series of MCL-1 inhibitors with high affinity and selectivity. J Med Chem58:2180-94 (2015) [PubMed] Article Bruncko, M; Wang, L; Sheppard, GS; Phillips, DC; Tahir, SK; Xue, J; Erickson, S; Fidanze, S; Fry, E; Hasvold, L; Jenkins, GJ; Jin, S; Judge, RA; Kovar, PJ; Madar, D; Nimmer, P; Park, C; Petros, AM; Rosenberg, SH; Smith, ML; Song, X; Sun, C; Tao, ZF; Wang, X; Xiao, Y; Zhang, H; Tse, C; Leverson, JD; Elmore, SW; Souers, AJ Structure-guided design of a series of MCL-1 inhibitors with high affinity and selectivity. J Med Chem58:2180-94 (2015) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Induced myeloid leukemia cell differentiation protein Mcl-1 | |||
| Name: | Induced myeloid leukemia cell differentiation protein Mcl-1 | ||
| Synonyms: | BCL2L3 | Bcl-2-like protein 3 | Bcl-2-like protein 3 (Mcl-1) | Bcl-2-related protein EAT/mcl1 | Bcl2-L-3 | Induced myeloid leukemia cell differentiation protein (Mcl-1) | MCL1 | MCL1_HUMAN | Mcl-1 | Myeloid Cell factor-1 (Mcl-1) | Myeloid cell leukemia sequence 1 (BCL2-related) | mcl1/EAT | ||
| Type: | Membrane; Single-pass membrane protein | ||
| Mol. Mass.: | 37332.87 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | Q07820 | ||
| Residue: | 350 | ||
| Sequence: |
| ||
| BDBM50078165 | |||
| n/a | |||
| Name | BDBM50078165 | ||
| Synonyms: | CHEMBL3417706 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C46H46N6O5 | ||
| Mol. Mass. | 762.8946 | ||
| SMILES | CC(=O)N1CCN(CC1)c1ccc(OCc2nn(C)c(C)c2-c2cccc3c(CCCOc4cccc5ccccc45)c(C(O)=O)n(Cc4cccnc4)c23)cc1 |(19.93,14.21,;19.31,15.27,;19.92,16.34,;17.77,15.26,;16.99,16.59,;15.45,16.59,;14.69,15.25,;15.46,13.92,;17,13.93,;13.15,15.25,;12.38,13.91,;10.84,13.91,;10.07,15.24,;8.52,15.23,;7.76,13.9,;6.22,13.89,;5.3,15.13,;3.84,14.63,;2.84,15.35,;3.86,13.09,;2.87,12.35,;5.32,12.66,;5.8,11.19,;7.33,10.86,;7.8,9.4,;6.76,8.24,;5.25,8.57,;4.01,7.7,;4.01,6.16,;2.67,5.39,;2.67,3.85,;1.33,3.08,;1.33,1.54,;2.66,.77,;2.66,-.77,;1.33,-1.54,;,-.77,;-1.33,-1.54,;-2.68,-.77,;-2.68,.77,;-1.33,1.54,;,.77,;2.78,8.56,;1.33,8.05,;.4,8.86,;1.09,6.84,;3.25,10.03,;2.34,11.27,;.8,11.1,;.19,9.69,;-1.34,9.51,;-2.26,10.75,;-1.64,12.16,;-.11,12.34,;4.78,10.03,;10.83,16.58,;12.37,16.58,)| | ||
| Structure | 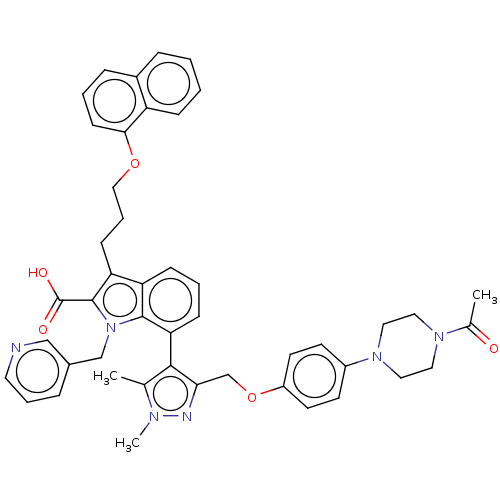
| ||