| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Voltage-dependent L-type calcium channel subunit alpha-1C |
|---|
| Ligand | BDBM50121975 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1479150 (CHEMBL3436126) |
|---|
| IC50 | 19820±n/a nM |
|---|
| Citation |  Wisniowska, B; Mendyk, A; Fijorek, K; Glinka, A; Polak, S Predictive model for L-type channel inhibition: multichannel block in QT prolongation risk assessment. J Appl Toxicol32:858-66 (2012) [PubMed] Article Wisniowska, B; Mendyk, A; Fijorek, K; Glinka, A; Polak, S Predictive model for L-type channel inhibition: multichannel block in QT prolongation risk assessment. J Appl Toxicol32:858-66 (2012) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Voltage-dependent L-type calcium channel subunit alpha-1C |
|---|
| Name: | Voltage-dependent L-type calcium channel subunit alpha-1C |
|---|
| Synonyms: | CAC1C_HUMAN | CACH2 | CACN2 | CACNA1C | CACNL1A1 | CCHL1A1 | Calcium channel (Type L) | Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle | L-type calcium channel alpha-1c/beta-2/alpha2delta-1 | Voltage-dependent L-type calcium channel subunit alpha-1C | Voltage-gated L-type calcium channel | Voltage-gated L-type calcium channel alpha-1C subunit | Voltage-gated calcium channel | Voltage-gated calcium channel subunit alpha Cav1.2 |
|---|
| Type: | Enzyme Catalytic Domain |
|---|
| Mol. Mass.: | 248979.79 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Calcium channel (Type L) 0 HUMAN::Q13936 |
|---|
| Residue: | 2221 |
|---|
| Sequence: | MVNENTRMYIPEENHQGSNYGSPRPAHANMNANAAAGLAPEHIPTPGAALSWQAAIDAAR
QAKLMGSAGNATISTVSSTQRKRQQYGKPKKQGSTTATRPPRALLCLTLKNPIRRACISI
VEWKPFEIIILLTIFANCVALAIYIPFPEDDSNATNSNLERVEYLFLIIFTVEAFLKVIA
YGLLFHPNAYLRNGWNLLDFIIVVVGLFSAILEQATKADGANALGGKGAGFDVKALRAFR
VLRPLRLVSGVPSLQVVLNSIIKAMVPLLHIALLVLFVIIIYAIIGLELFMGKMHKTCYN
QEGIADVPAEDDPSPCALETGHGRQCQNGTVCKPGWDGPKHGITNFDNFAFAMLTVFQCI
TMEGWTDVLYWVNDAVGRDWPWIYFVTLIIIGSFFVLNLVLGVLSGEFSKEREKAKARGD
FQKLREKQQLEEDLKGYLDWITQAEDIDPENEDEGMDEEKPRNMSMPTSETESVNTENVA
GGDIEGENCGARLAHRISKSKFSRYWRRWNRFCRRKCRAAVKSNVFYWLVIFLVFLNTLT
IASEHYNQPNWLTEVQDTANKALLALFTAEMLLKMYSLGLQAYFVSLFNRFDCFVVCGGI
LETILVETKIMSPLGISVLRCVRLLRIFKITRYWNSLSNLVASLLNSVRSIASLLLLLFL
FIIIFSLLGMQLFGGKFNFDEMQTRRSTFDNFPQSLLTVFQILTGEDWNSVMYDGIMAYG
GPSFPGMLVCIYFIILFICGNYILLNVFLAIAVDNLADAESLTSAQKEEEEEKERKKLAR
TASPEKKQELVEKPAVGESKEEKIELKSITADGESPPATKINMDDLQPNENEDKSPYPNP
ETTGEEDEEEPEMPVGPRPRPLSELHLKEKAVPMPEASAFFIFSSNNRFRLQCHRIVNDT
IFTNLILFFILLSSISLAAEDPVQHTSFRNHILFYFDIVFTTIFTIEIALKILGNADYVF
TSIFTLEIILKMTAYGAFLHKGSFCRNYFNILDLLVVSVSLISFGIQSSAINVVKILRVL
RVLRPLRAINRAKGLKHVVQCVFVAIRTIGNIVIVTTLLQFMFACIGVQLFKGKLYTCSD
SSKQTEAECKGNYITYKDGEVDHPIIQPRSWENSKFDFDNVLAAMMALFTVSTFEGWPEL
LYRSIDSHTEDKGPIYNYRVEISIFFIIYIIIIAFFMMNIFVGFVIVTFQEQGEQEYKNC
ELDKNQRQCVEYALKARPLRRYIPKNQHQYKVWYVVNSTYFEYLMFVLILLNTICLAMQH
YGQSCLFKIAMNILNMLFTGLFTVEMILKLIAFKPKGYFSDPWNVFDFLIVIGSIIDVIL
SETNHYFCDAWNTFDALIVVGSIVDIAITEVNPAEHTQCSPSMNAEENSRISITFFRLFR
VMRLVKLLSRGEGIRTLLWTFIKSFQALPYVALLIVMLFFIYAVIGMQVFGKIALNDTTE
INRNNNFQTFPQAVLLLFRCATGEAWQDIMLACMPGKKCAPESEPSNSTEGETPCGSSFA
VFYFISFYMLCAFLIINLFVAVIMDNFDYLTRDWSILGPHHLDEFKRIWAEYDPEAKGRI
KHLDVVTLLRRIQPPLGFGKLCPHRVACKRLVSMNMPLNSDGTVMFNATLFALVRTALRI
KTEGNLEQANEELRAIIKKIWKRTSMKLLDQVVPPAGDDEVTVGKFYATFLIQEYFRKFK
KRKEQGLVGKPSQRNALSLQAGLRTLHDIGPEIRRAISGDLTAEEELDKAMKEAVSAASE
DDIFRRAGGLFGNHVSYYQSDGRSAFPQTFTTQRPLHINKAGSSQGDTESPSHEKLVDST
FTPSSYSSTGSNANINNANNTALGRLPRPAGYPSTVSTVEGHGPPLSPAIRVQEVAWKLS
SNRERHVPMCEDLELRRDSGSAGTQAHCLLLRKANPSRCHSRESQAAMAGQEETSQDETY
EVKMNHDTEACSEPSLLSTEMLSYQDDENRQLTLPEEDKRDIRQSPKRGFLRSASLGRRA
SFHLECLKRQKDRGGDISQKTVLPLHLVHHQALAVAGLSPLLQRSHSPASFPRPFATPPA
TPGSRGWPPQPVPTLRLEGVESSEKLNSSFPSIHCGSWAETTPGGGGSSAARRVRPVSLM
VPSQAGAPGRQFHGSASSLVEAVLISEGLGQFAQDPKFIEVTTQELADACDMTIEEMESA
ADNILSGGAPQSPNGALLPFVNCRDAGQDRAGGEEDAGCVRARGRPSEEELQDSRVYVSS
L
|
|
|
|---|
| BDBM50121975 |
|---|
| n/a |
|---|
| Name | BDBM50121975 |
|---|
| Synonyms: | (6-Methoxy-quinolin-4-yl)-(5-vinyl-1-aza-bicyclo[2.2.2]oct-2-yl)-methanol | CARDIOQUIN | CHEMBL1294 | CIN-QUIN | DURAQUIN | QUINACT | QUINAGLUTE | QUINALAN | QUINATIME | QUINIDEX | QUINIDINE | QUINORA | US9402878, Quinidine |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C20H24N2O2 |
|---|
| Mol. Mass. | 324.4168 |
|---|
| SMILES | COc1ccc2nccc([C@H](O)[C@H]3C[C@@H]4CCN3C[C@@H]4C=C)c2c1 |r,THB:20:19:12.13:16.15,10:12:18.19:16.15| |
|---|
| Structure | 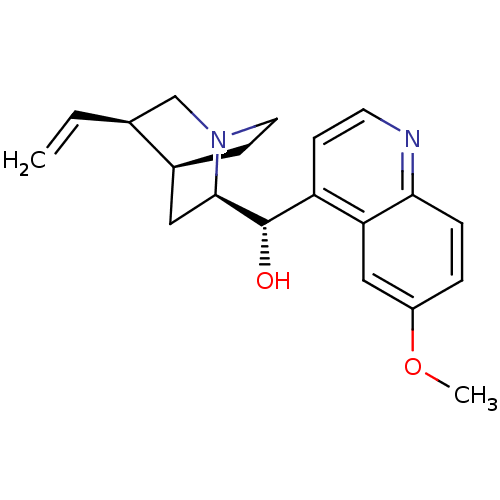
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Wisniowska, B; Mendyk, A; Fijorek, K; Glinka, A; Polak, S Predictive model for L-type channel inhibition: multichannel block in QT prolongation risk assessment. J Appl Toxicol32:858-66 (2012) [PubMed] Article
Wisniowska, B; Mendyk, A; Fijorek, K; Glinka, A; Polak, S Predictive model for L-type channel inhibition: multichannel block in QT prolongation risk assessment. J Appl Toxicol32:858-66 (2012) [PubMed] Article