| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Lysine-specific demethylase 4E |
|---|
| Ligand | BDBM50153322 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1562434 (CHEMBL3778170) |
|---|
| IC50 | 40±n/a nM |
|---|
| Citation |  Westaway, SM; Preston, AG; Barker, MD; Brown, F; Brown, JA; Campbell, M; Chung, CW; Diallo, H; Douault, C; Drewes, G; Eagle, R; Gordon, L; Haslam, C; Hayhow, TG; Humphreys, PG; Joberty, G; Katso, R; Kruidenier, L; Leveridge, M; Liddle, J; Mosley, J; Muelbaier, M; Randle, R; Rioja, I; Rueger, A; Seal, GA; Sheppard, RJ; Singh, O; Taylor, J; Thomas, P; Thomson, D; Wilson, DM; Lee, K; Prinjha, RK Cell Penetrant Inhibitors of the KDM4 and KDM5 Families of Histone Lysine Demethylases. 1. 3-Amino-4-pyridine Carboxylate Derivatives. J Med Chem59:1357-69 (2016) [PubMed] Article Westaway, SM; Preston, AG; Barker, MD; Brown, F; Brown, JA; Campbell, M; Chung, CW; Diallo, H; Douault, C; Drewes, G; Eagle, R; Gordon, L; Haslam, C; Hayhow, TG; Humphreys, PG; Joberty, G; Katso, R; Kruidenier, L; Leveridge, M; Liddle, J; Mosley, J; Muelbaier, M; Randle, R; Rioja, I; Rueger, A; Seal, GA; Sheppard, RJ; Singh, O; Taylor, J; Thomas, P; Thomson, D; Wilson, DM; Lee, K; Prinjha, RK Cell Penetrant Inhibitors of the KDM4 and KDM5 Families of Histone Lysine Demethylases. 1. 3-Amino-4-pyridine Carboxylate Derivatives. J Med Chem59:1357-69 (2016) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Lysine-specific demethylase 4E |
|---|
| Name: | Lysine-specific demethylase 4E |
|---|
| Synonyms: | 2-OG-Dependent Histone Demethylase JMJD2E | Histone Lysine Demethylase | Jumonji domain-containing protein 2E (JMJD2E) | KDM4D-like protein | KDM4DL | KDM4E | KDM4E_HUMAN | Lysine-specific demethylase 4D-like | Lysine-specific demethylase 4E | Lysine-specific demethylase 4E (KDM4E) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 56815.71 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | B2RXH2 |
|---|
| Residue: | 506 |
|---|
| Sequence: | MKSVHSSPQNTSHTIMTFYPTMEEFADFNTYVAYMESQGAHQAGLAKVIPPKEWKARQMY
DDIEDILIATPLQQVTSGQGGVFTQYHKKKKAMRVGQYRRLANSKKYQTPPHQNFADLEQ
RYWKSHPGNPPIYGADISGSLFEESTKQWNLGHLGTILDLLEQECGVVIEGVNTPYLYFG
MWKTTFAWHTEDMDLYSINYLHFGEPKTWYVVPPEHGQHLERLARELFPDISRGCEAFLR
HKVALISPTVLKENGIPFNCMTQEAGEFMVTFPYGYHAGFNHGFNCAEAINFATPRWIDY
GKMASQCSCGESTVTFSMDPFVRIVQPESYELWKHRQDLAIVEHTEPRVAESQELSNWRD
DIVLRRAALGLRLLPNLTAQCPTQPVSSGHCYNPKGCGTDAVPGSAFQSSAYHTQTQSLT
LGMSARVLLPSTGSWGSGRGRGRGQGQGRGCSRGRGHGCCTRELGTEEPTVQPASKRRLL
MGTRSRAQGHRPQLPLANDLMTNLSL
|
|
|
|---|
| BDBM50153322 |
|---|
| n/a |
|---|
| Name | BDBM50153322 |
|---|
| Synonyms: | CHEMBL3775612 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C13H12N2O2 |
|---|
| Mol. Mass. | 228.2466 |
|---|
| SMILES | OC(=O)c1ccncc1NCc1ccccc1 |
|---|
| Structure | 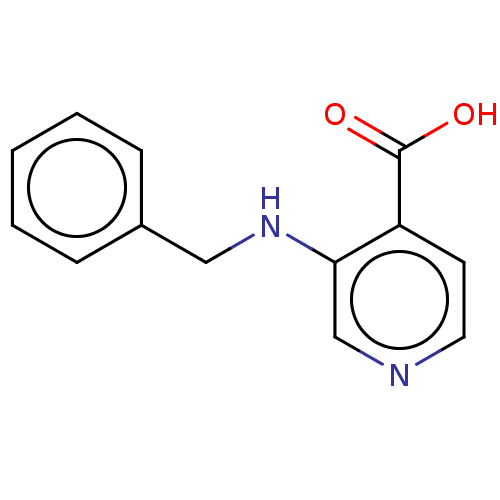
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Westaway, SM; Preston, AG; Barker, MD; Brown, F; Brown, JA; Campbell, M; Chung, CW; Diallo, H; Douault, C; Drewes, G; Eagle, R; Gordon, L; Haslam, C; Hayhow, TG; Humphreys, PG; Joberty, G; Katso, R; Kruidenier, L; Leveridge, M; Liddle, J; Mosley, J; Muelbaier, M; Randle, R; Rioja, I; Rueger, A; Seal, GA; Sheppard, RJ; Singh, O; Taylor, J; Thomas, P; Thomson, D; Wilson, DM; Lee, K; Prinjha, RK Cell Penetrant Inhibitors of the KDM4 and KDM5 Families of Histone Lysine Demethylases. 1. 3-Amino-4-pyridine Carboxylate Derivatives. J Med Chem59:1357-69 (2016) [PubMed] Article
Westaway, SM; Preston, AG; Barker, MD; Brown, F; Brown, JA; Campbell, M; Chung, CW; Diallo, H; Douault, C; Drewes, G; Eagle, R; Gordon, L; Haslam, C; Hayhow, TG; Humphreys, PG; Joberty, G; Katso, R; Kruidenier, L; Leveridge, M; Liddle, J; Mosley, J; Muelbaier, M; Randle, R; Rioja, I; Rueger, A; Seal, GA; Sheppard, RJ; Singh, O; Taylor, J; Thomas, P; Thomson, D; Wilson, DM; Lee, K; Prinjha, RK Cell Penetrant Inhibitors of the KDM4 and KDM5 Families of Histone Lysine Demethylases. 1. 3-Amino-4-pyridine Carboxylate Derivatives. J Med Chem59:1357-69 (2016) [PubMed] Article